sliceMetrics
Description
sliceMetrics computes group metrics on slices of data using a
trained machine learning model (that is, a binary classifier or a regression model). You can
use the metrics to determine whether the model performs similarly across the slices of
data.
After creating a sliceMetrics object, use the report function to
generate a slice metrics report, or use the plot function to
create a bar graph of a slice metric. Some metrics directly compare data slices to their
complements. The complement of a data slice consists of all observations that are not in the
data slice.
Creation
Syntax
Description
Input Arguments
Machine learning model on which to calculate metrics, specified as a binary classification model object or a regression model object in one of the following tables.
| Model | Full or Compact Model Object |
|---|---|
| Discriminant analysis classifier | ClassificationDiscriminant, CompactClassificationDiscriminant |
| Ensemble of learners for classification | ClassificationEnsemble, CompactClassificationEnsemble,
ClassificationBaggedEnsemble |
| Generalized additive model (GAM) | ClassificationGAM, CompactClassificationGAM |
| Gaussian kernel classification model using random feature expansion | ClassificationKernel |
| k-nearest neighbor classifier | ClassificationKNN |
| Linear classification model | ClassificationLinear |
| Naive Bayes model | ClassificationNaiveBayes, CompactClassificationNaiveBayes |
| Neural network classifier | ClassificationNeuralNetwork, CompactClassificationNeuralNetwork |
| Support vector machine (SVM) for binary classification | ClassificationSVM, CompactClassificationSVM |
| Decision tree for binary classification | ClassificationTree, CompactClassificationTree |
| Model | Full or Compact Model Object |
|---|---|
| Ensemble of regression models | RegressionEnsemble, RegressionBaggedEnsemble, CompactRegressionEnsemble |
| Generalized additive model (GAM) | RegressionGAM, CompactRegressionGAM |
| Gaussian process regression | RegressionGP, CompactRegressionGP |
| Gaussian kernel regression model using random feature expansion | RegressionKernel |
| Linear regression for high-dimensional data | RegressionLinear |
| Neural network regression model | RegressionNeuralNetwork, CompactRegressionNeuralNetwork |
| Support vector machine (SVM) regression | RegressionSVM, CompactRegressionSVM |
| Regression tree | RegressionTree, CompactRegressionTree |
If Mdl is a model object that does not contain predictor data
(for example, a compact model), you must provide the input argument X.
Variables to slice or indices of a data slice, specified as one of the values in the following table.
| Description | Value |
|---|---|
| One or two variables to slice |
|
| Indices of a data slice | Logical vector of length equal to the number of observations in the data
X, where true values indicate
observations in the data slice |
Data Types: single | double | logical | char | string | cell
Data to use for computing metrics, specified as a numeric matrix or a table. Each
row of X corresponds to one observation, and each column
corresponds to one variable.
For a numeric matrix:
The variables that make up the columns of
Xmust have the same order as the predictor variables that trainedMdl(that is,Mdl.PredictorNames).If you trained
Mdlusing a table, thenXcan be a numeric matrix if the table contains all numeric predictor variables.
For a table:
If you trained
Mdlusing a table (for example,Tbl), then all predictor variables inXmust have the same variable names and data types as those inTbl. However, the column order ofXdoes not need to correspond to the column order ofTbl.If you trained
Mdlusing a numeric matrix, then the predictor names inMdl.PredictorNamesmust be the same as the corresponding predictor variable names inX. To specify predictor names during training, use thePredictorNamesname-value argument. All predictor variables inXmust be numeric vectors.Xcan contain additional variables.sliceMetricsdoes not support multicolumn variables or cell arrays other than cell arrays of character vectors.
If X does not include the response data, specify Y.
Data Types: single | double | table
Response data to use for computing metrics, specified as a numeric, categorical,
or logical vector; a character or string array; or a cell array of character vectors.
Y must have the same number of observations as X, and the
data type of Y must match the data type of the response variable
used to train Mdl.
Data Types: single | double | logical | char | string | cell | categorical
Number of bins for the numeric predictors listed in slices,
specified as a positive integer scalar or two-element positive integer vector. If you
specify to slice two numeric predictors and numbins is a scalar,
the function partitions each predictor's values into numbins
bins. For more information, see Binning of Predictors.
Data Types: single | double
Properties
This property is read-only.
Slice indicator for each observation in Mdl.X or
X, returned as a numeric vector. For example, if you use
X to create the object sliceResults, then
X(sliceResults.SliceIndex==i,:) returns the observations in slice
i.
Data Types: double
This property is read-only.
Variables used to create the data slices, returned as a string array. If you specify
data slice indices when you create the object, then the
SliceVariables value is "custom". Otherwise,
SliceVariables contains the names of predictors listed in
Mdl.PredictorNames.
Data Types: string
Examples
Train a binary classifier on numeric data. Use sliceMetrics to slice the training data according to one of the predictors. Evaluate the accuracy of the model predictions on the data slices.
Load the sample file fisheriris.csv, which contains iris data including sepal length, sepal width, petal width, and species type. Read the file into a table, and then convert the Species variable into a categorical variable. Display the first eight observations in the table.
fisheriris = readtable("fisheriris.csv");
fisheriris.Species = categorical(fisheriris.Species);
head(fisheriris) SepalLength SepalWidth PetalLength PetalWidth Species
___________ __________ ___________ __________ _______
5.1 3.5 1.4 0.2 setosa
4.9 3 1.4 0.2 setosa
4.7 3.2 1.3 0.2 setosa
4.6 3.1 1.5 0.2 setosa
5 3.6 1.4 0.2 setosa
5.4 3.9 1.7 0.4 setosa
4.6 3.4 1.4 0.3 setosa
5 3.4 1.5 0.2 setosa
Separate the data for two of the iris species: versicolor and virginica.
versicolorData = fisheriris(fisheriris.Species=="versicolor",:); virginicaData = fisheriris(fisheriris.Species=="virginica",:); trainingData = [versicolorData;virginicaData];
Train a binary tree classifier on the versicolor and virginica data.
Mdl = fitctree(trainingData,"Species")Mdl =
ClassificationTree
PredictorNames: {'SepalLength' 'SepalWidth' 'PetalLength' 'PetalWidth'}
ResponseName: 'Species'
CategoricalPredictors: []
ClassNames: [versicolor virginica]
ScoreTransform: 'none'
NumObservations: 100
Properties, Methods
Mdl is a ClassificationTree model object trained on 100 observations.
Compute metrics on the training data slices determined by petal length. Because PetalLength is a numeric predictor, sliceMetrics creates data slices by binning the petal length values of observations in Mdl.X. Data slices always partition the data.
sliceResults = sliceMetrics(Mdl,"PetalLength")sliceResults =
sliceMetrics evaluated on PetalLength slices:
PetalLength NumObservations Accuracy OddsRatio PValue EffectSize
___________ _______________ ________ _________ ________ __________
[3, 3.65) 6 1 0 0.083246 -0.031915
[3.65, 4.3) 17 1 0 0.083241 -0.036145
[4.3, 4.95) 31 0.96774 1.1167 0.93193 0.0032726
[4.95, 5.6) 21 0.90476 8.2105 0.23003 0.08258
[5.6, 6.25) 19 1 0 0.08324 -0.037037
[6.25, 6.9] 6 1 0 0.083246 -0.031915
Properties, Methods
In this example, sliceMetrics creates six data slices. The accuracy for each data slice is quite high (over 90%).
Visualize the accuracy for each slice and its complement in Mdl.X.
plot(sliceResults)
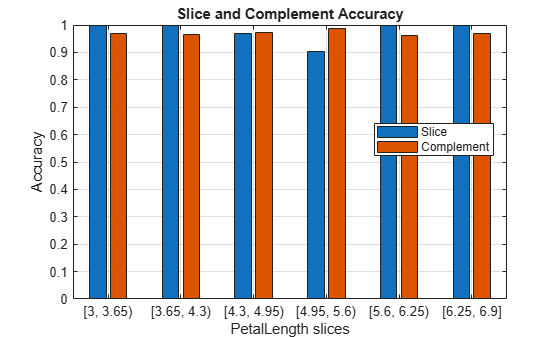
In general, the accuracy for each slice is similar to the accuracy for its slice complement. However, the observations with petal lengths in the range [4.95,5.6) have a slightly lower percentage of correct classifications than all other observations.
Train a regression model using a mix of numeric and categorical data. Use sliceMetrics to compute metrics on a specified data slice of interest.
Load the carbig data set, which contains measurements of cars made in the 1970s and early 1980s. Convert the Origin data to a categorical variable, and combine the variable with a subset of the other measurements into a table.
load carbig Origin = categorical(cellstr(Origin)); cars = table(Acceleration,Cylinders,Displacement,Horsepower, ... Origin,Weight,MPG);
Remove observations with missing values from the cars table. Then, display the first eight observations in the table.
cars = rmmissing(cars); head(cars)
Acceleration Cylinders Displacement Horsepower Origin Weight MPG
____________ _________ ____________ __________ ______ ______ ___
12 8 307 130 USA 3504 18
11.5 8 350 165 USA 3693 15
11 8 318 150 USA 3436 18
12 8 304 150 USA 3433 16
10.5 8 302 140 USA 3449 17
10 8 429 198 USA 4341 15
9 8 454 220 USA 4354 14
8.5 8 440 215 USA 4312 14
Partition the data into training data and test data. Reserve approximately 50% of the observations for computing slice metrics, and use the rest of the observations for model training.
rng(0,"twister") % For reproducibility cv = cvpartition(length(cars.MPG),Holdout=0.5); trainingCars = cars(training(cv),:); testCars = cars(test(cv),:);
Train a Gaussian process regression model using the training data. Standardize the numeric predictors before fitting the model.
Mdl = fitrgp(trainingCars,"MPG",Standardize=true)Mdl =
RegressionGP
PredictorNames: {'Acceleration' 'Cylinders' 'Displacement' 'Horsepower' 'Origin' 'Weight'}
ResponseName: 'MPG'
CategoricalPredictors: 5
ResponseTransform: 'none'
NumObservations: 196
KernelFunction: 'SquaredExponential'
KernelInformation: [1×1 struct]
BasisFunction: 'Constant'
Beta: 25.8166
Sigma: 3.9677
PredictorLocation: [11×1 double]
PredictorScale: [11×1 double]
Alpha: [196×1 double]
ActiveSetVectors: [196×11 double]
PredictMethod: 'Exact'
ActiveSetSize: 196
FitMethod: 'Exact'
ActiveSetMethod: 'Random'
IsActiveSetVector: [196×1 logical]
LogLikelihood: -566.1334
ActiveSetHistory: []
BCDInformation: []
Properties, Methods
Mdl is a RegressionGP model object trained on a mix of numeric and categorical predictors.
Create a data slice of the test set cars manufactured in the USA with an acceleration value of 15 or more.
testDataSliceIndex = testCars.Origin=="USA" & testCars.Acceleration >= 15;Evaluate the regression model on the custom data slice using the sliceMetrics function. Use the report function to display the mean squared error (MSE), a two-sample t-statistic, and a two-sample p-value for the custom data slice (true) and its complement (false).
sliceResults = sliceMetrics(Mdl,testCars,testDataSliceIndex); metricsTbl = report(sliceResults,Metrics=["mse","tstat","pvalue"])
metricsTbl=2×5 table
custom NumObservations Error TStatistic PValue
______ _______________ ______ __________ ________
true 61 9.624 -1.8661 0.063547
false 135 16.036 1.8661 0.063547
The MSE is smaller for the custom data slice than for the remaining test set observations. However, the t-statistic and p-value for Welch's t-test indicate that the mean of the squared errors is not statistically different at the 5% significance level between the slice and its complement.
Train a regression model on a mix of numeric and categorical data. Use sliceMetrics to slice the test data according to two predictors. Compute the mean squared error for the data slices. To improve the general model performance across the data slices, generate synthetic observations and use them to retrain the model.
Load the carbig data set, which contains measurements of cars made in the 1970s and early 1980s. Bin the Model_Year data to form a categorical variable, and combine the variable with a subset of the other measurements into a table. Remove observations with missing values from the table. Then, display the first eight observations in the table.
load carbig ModelDecade = discretize(Model_Year,[70 80 89], ... "categorical",["70s","80s"]); ModelDecade = categorical(ModelDecade,Ordinal=false); cars = table(Acceleration,Displacement,Horsepower, ... ModelDecade,Weight,MPG); cars = rmmissing(cars); head(cars)
Acceleration Displacement Horsepower ModelDecade Weight MPG
____________ ____________ __________ ___________ ______ ___
12 307 130 70s 3504 18
11.5 350 165 70s 3693 15
11 318 150 70s 3436 18
12 304 150 70s 3433 16
10.5 302 140 70s 3449 17
10 429 198 70s 4341 15
9 454 220 70s 4354 14
8.5 440 215 70s 4312 14
Partition the data into training data and test data. Reserve approximately 50% of the observations for computing slice metrics, and use the rest of the observations for model training.
rng(0,"twister") % For reproducibility of partition cv = cvpartition(length(cars.MPG),Holdout=0.5); trainingCars = cars(training(cv),:); testCars = cars(test(cv),:);
Train a regression tree model using the training data. Then, compute metrics on the test data slices determined by the decade of manufacture and the weight of the car. Partition the numeric Weight values into three bins. Because ModelDecade is a categorical variable with two categories, sliceMetrics creates six data slices.
Mdl = fitrtree(trainingCars,"MPG"); sliceResults = sliceMetrics(Mdl,testCars,["ModelDecade","Weight"], ... NumBins=3)
sliceResults =
sliceMetrics evaluated on ModelDecade and Weight slices:
ModelDecade Weight NumObservations Error TStatistic PValue EffectSize
___________ ________________ _______________ ______ __________ __________ __________
70s [1649, 2812.7) 64 18.392 0.23812 0.81224 1.3097
70s [2812.7, 3976.3) 53 6.6804 -4.3456 2.3745e-05 -14.843
70s [3976.3, 5140] 36 5.6089 -4.7484 4.0884e-06 -14.578
80s [1649, 2812.7) 32 30.874 2.2891 0.02727 15.972
80s [2812.7, 3976.3) 11 64.626 2.4672 0.032784 49.918
80s [3976.3, 5140] 0 NaN NaN NaN NaN
Properties, Methods
For the cars made in the 80s with a weight in the range [2812.7,3976.3) (that is, the observations in slice 5), the mean squared error (MSE) is much higher than the MSE for the cars in the other data slices.
Generate 500 synthetic observations using the synthesizeTabularData function. By default, the function uses a binning technique to learn the distribution of the variables in trainingCars before synthesizing the data.
rng(10,"twister") % For reproducibility of data generation syntheticCars = synthesizeTabularData(trainingCars,500);
Combine the training observations with the synthetic observations. Use the combined data to retrain the regression tree model.
newTrainingCars = [trainingCars;syntheticCars];
newMdl = fitrtree(newTrainingCars,"MPG");Compute metrics on the same test data slices using the retrained model.
newSliceResults = sliceMetrics(newMdl,testCars,["ModelDecade","Weight"], ... NumBins=3)
newSliceResults =
sliceMetrics evaluated on ModelDecade and Weight slices:
ModelDecade Weight NumObservations Error TStatistic PValue EffectSize
___________ ________________ _______________ ______ __________ __________ __________
70s [1649, 2812.7) 64 20.986 1.5012 0.13736 8.5476
70s [2812.7, 3976.3) 53 9.9331 -2.0166 0.045209 -7.2595
70s [3976.3, 5140] 36 5.4484 -4.3062 2.6575e-05 -11.982
80s [1649, 2812.7) 32 20.422 1.0664 0.29206 6.206
80s [2812.7, 3976.3) 11 24.162 1.0643 0.30927 9.463
80s [3976.3, 5140] 0 NaN NaN NaN NaN
Properties, Methods
Visually compare the MSE values for the data slices.
tiledlayout(1,2) nexttile plot(sliceResults) ylim([0 70]) xticklabels(1:6) legend(Location="northwest") nexttile plot(newSliceResults) ylim([0 70]) xticklabels(1:6) legend(Location="northwest")
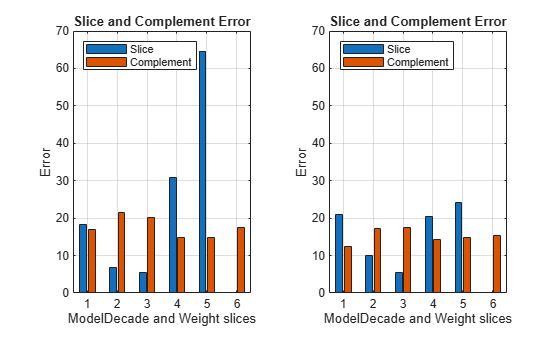
The MSE for slice 5 is lower for the model trained with the training and synthetic observations (newMdl) than for the model trained with only the training data (Mdl).
Tips
If your model does not perform well across all data slices, you can try the following:
Retrain your model using different observation weights.
Collect new data or generate synthetic data before retraining your model. For an example, see Improve Performance on Data Slices Using Synthetic Data.
Algorithms
To create data slices, sliceMetrics bins each numeric variable
listed in slices as follows, where m is the number
of observations in the data after the software removes observations with missing response values:
Equiprobable binning with
ceil(2*m^(2/5))bins when the data contains nonfinite values and you do not specify the number of bins (numbins)Equiprobable binning when the data contains nonfinite values and you specify the number of bins (
numbins)Equal-width binning when the data contains finite values only and you specify the number of bins (
numbins)Equal-width binning with
ceil((2*m)^(1/3))bins otherwise
If you create data slices using a categorical predictor (that is, a predictor variable
whose index is in Mdl.CategoricalPredictors), the function uses the
categories of the predictor to partition the data.
References
[1] Chung, Yeounoh, Tim Kraska, Neoklis Polyzotis, Ki Hyun Tae, and Steven Euijong Whang. “Automated Data Slicing for Model Validation: A Big Data - AI Integration Approach.” IEEE Transactions on Knowledge and Data Engineering 32, no. 12 (2020): 2284–96.
Version History
Introduced in R2026a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Web サイトの選択
Web サイトを選択すると、翻訳されたコンテンツにアクセスし、地域のイベントやサービスを確認できます。現在の位置情報に基づき、次のサイトの選択を推奨します:
また、以下のリストから Web サイトを選択することもできます。
最適なサイトパフォーマンスの取得方法
中国のサイト (中国語または英語) を選択することで、最適なサイトパフォーマンスが得られます。その他の国の MathWorks のサイトは、お客様の地域からのアクセスが最適化されていません。
南北アメリカ
- América Latina (Español)
- Canada (English)
- United States (English)
ヨーロッパ
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)