timeSeriesIforestAD
Create a machine learning isolation forest anomaly detector model for time series data
Since R2026a
Syntax
Description
Add-On Required: This feature requires the Time Series Anomaly Detection for MATLAB add-on.
timeSeriesIforestAD creates an anomaly detector based on the
machine learning Isolated Forest algorithm. This algorithm is detects
anomalies by isolating anomalous data from normal data using an ensemble of isolation
trees.
For more information on the model that timeSeriesIforestAD creates,
see TimeSeriesIForestDetector.
detector = timeSeriesIforestAD(numChannels)TimeseriesIForestDetector model for time series data with
numChannels input channels.
detector = timeSeriesIforestAD(
sets additional options using one or more name-value arguments.numChannels,Name=Value)
For example, detector = timeSeriesIforestAD(3,NumLearners=120)
creates a an isolation forest with three input channels and 120 trees. detector =
timeSeriesIforestAD(3,DetectionWindowLength=20) creates a detector model with a
detection window length of 20.
Examples
Load the file sineWaveAnomalyData.mat, which contains two sets of synthetic 3-channel sinusoidal signals.
sineWaveNormal contains 10 sinusoids of stable frequency and amplitude. Each signal has a series of small-amplitude impact-like imperfections. The signals have different lengths and initial phases. sineWaveAbnormal contains 3 sinusoids that contain the same normal data as sineWaveNormal, but that also include anomalous data.
load sineWaveAnomalyData.mat sineWaveNormal sineWaveAbnormal
Plot input signals
Plot all 3 channels of the first three anomalous signals.
s1 = 3; tiledlayout("vertical") ax = zeros(s1,1); for kj = 1:s1 ax(kj) = nexttile; plot(sineWaveAbnormal{kj}) title("Anomalous Signals") end
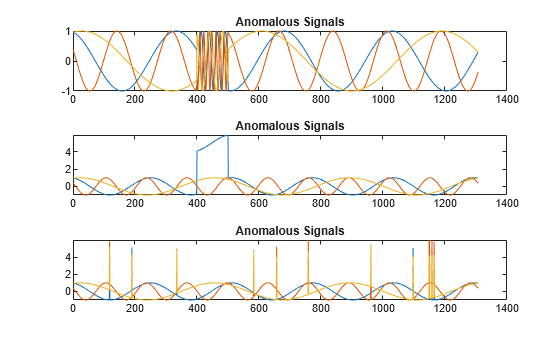
sineWaveAbnormal contains three signals, all of the same length. Each signal in the set has one or more anomalies.
All channels of the first signal have an abrupt change in frequency that lasts for a finite time.
The second signal has a finite-duration amplitude change in one of its channels.
The third signal has spikes at random times in all channels.
Create Detector
Use the timeseriesIforestAD detector to create an Isolated Forest detector with 3 channels and default options.
detector_tsif = timeSeriesIforestAD(3)
detector_tsif =
TimeSeriesIForestDetector with properties:
NumLearners: 100
NumObservationsPerLearner: []
NumChannels: 3
IsTrained: 0
WindowLength: 10
TrainingStride: 1
DetectionStride: 10
Threshold: []
ThresholdMethod: "kSigma"
ThresholdParameter: 3
ThresholdFunction: []
Normalization: "zscore"
FeatureExtraction: 1
Train detector using the normal data.
detector_tsif = train(detector_tsif,sineWaveNormal);
View the threshold that train computes and saves within detector_tsif. This computed value is influenced by random factors, such as which subsets of the data are used for training, and can change somewhat for different training sessions and different machines.
thresh = detector_tsif.Threshold
thresh = 0.5867
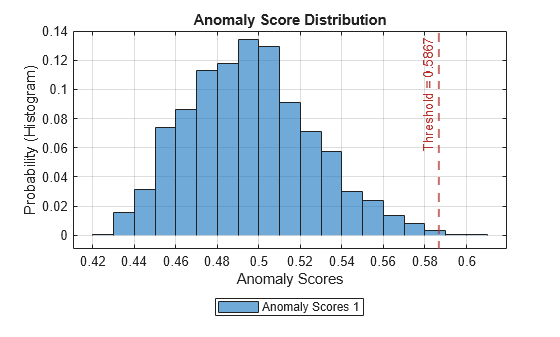
Plot the histogram of the anomaly scores for the normal data. Each score is calculated over a single detection window. The threshold, plotted as a vertical line, does not always completely bound the scores.
plotHistogram(detector_tsif,sineWaveNormal);
Use Detector to Identify Anomalies
Use the detect function to determine the anomaly scores for the anomalous data. Then, plot the anomaly scores.
results = detect(detector_tsif, sineWaveAbnormal)
results=3×1 cell array
{130×3 table}
{130×3 table}
{130×3 table}
results is a cell array that contains three tables, one table for each signal. Each cell table contains three variables: WindowLabel, WindowAnomalyScore, and WindowStartIndices. Confirm the table variable names.
varnames = results{1}.Properties.VariableNamesvarnames = 1×3 cell
{'Labels'} {'AnomalyScores'} {'StartIndices'}
Plot Anomaly Score Distributions
Plot a histogram that shows the anomaly scores for both sets of data together, along with the threshold, for comparison.
plotHistogram(detector_tsif,sineWaveNormal,sineWaveAbnormal)
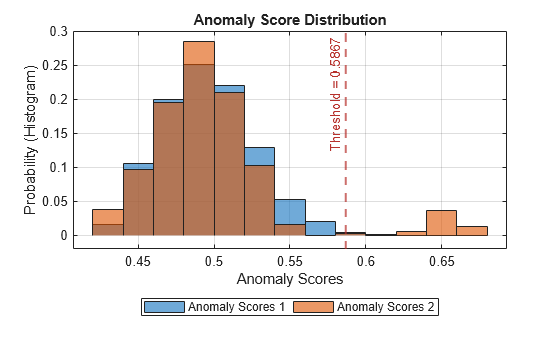
The histogram uses different colors for the normal (Data 1) and anomalous (Data 2) data. Both types of data appear to the left of the threshold. To the right of threshold, Data 2 is prevalent.
Plot the detected anomalies of the third abnormal signal set.
plot(detector_tsif,sineWaveAbnormal{3})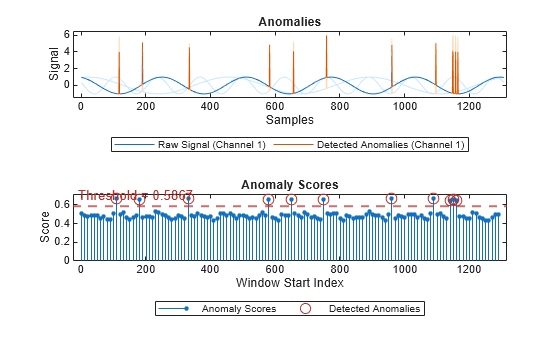
The top plot shows an overlay of red where the anomalies occur. The bottom plot shows how effective the threshold is at dividing the normal from the abnormal scores for Signal set 3.
Input Arguments
Number of input channels in each time series, specified as a positive integer. All time series inputs must have the same number of channels.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: detector = timeSeriesIforestAD(3,DetectionWindowLength=20)
sets the length of the detection window to 20.
Window
Window length of each time series segment, specified as a positive integer.
Stride length of sliding window in training stage, specified as a positive integer.
Stride length of sliding window in detection stage, specified as a positive
integer. DetectionStride controls the number of overlapped
samples. Increasing the amount of overlap provides higher resolution, but at higher
computational cost.
If you do not specify DetectionStride, the software sets the
stride length to the value of WindowLength to create
non-overlapping windows.
Feature Extraction
Enable feature extraction option, specified as 1(true) or
0(false).
When
FeatureExtractionis enabled, the software extracts statistical features to use during training. Extracted features are generally cleaner and more meaningful than raw data, but the extraction process can be computationally expensive.When
FeatureExtractionis disabled, the training process uses only raw data. Raw data is likely to be noisier and include random disturbances. However, if you have a relatively small number of signals and they already represent useful features, then the computational costFeatureExtractionmay not be worth the benefit.
Threshold
Method to compute the detection threshold, specified as one of these values, each of which correspond to what the detection threshold is based on:
"kSigma"— Standard deviation of the normalized anomaly scores. The parameter k determines the threshold within the standard deviation levels that identifies an anomaly. The value of k is specified byThresholdParameter."contaminationFraction"— Percentage of anomalies within a specified fraction of windows, measured over the entire training set. The fraction value is specified byThresholdParameter."max"— Maximum window loss measured over the entire training data set and multiplied byThresholdParameter"mean"— Mean window loss measured over the entire training data set and multiplied byThresholdParameter"median"— Median window loss measured over the entire training data set and multiplied byThresholdParameter"manual"— Manual detection threshold value based onThreshold."customFunction"— Custom detection threshold method based onThresholdFunction.
If you specify ThresholdMethod, you can also specify
ThresholdParameter, Threshold, or ThresholdParameter. The
available threshold parameter depends on the specified detection method.
Anomaly score used to detect anomalies, specified as a positive scalar. The source
of the Threshold value depends on the setting of ThresholdMethod.
If
ThresholdMethodis"manual", you set the value.If
ThresholdMethodis"customFunction", the function specified inThresholdFunctioncomputes the value.For other values of
ThresholdMethod, useThresholdParameterto specify the detection threshold.
Parameter used for determining the detection threshold, specified as a numeric scalar.
The way you specify ThresholdParameter depends on the
specified value for ThresholdMethod. The following list describes
the specification of ThresholdParameter for each possible value
of ThresholdMethod
"kSigma"— SpecifyThresholdParameteras a positive numeric scalar. If you do not specifyThresholdParameter, the detector sets the threshold to 3."contaminationFraction"— SpecifyThresholdParameteras a as a nonnegative scalar less than 0.5. For example, if you specify"contaminationFraction"as0.05, then the threshold is set to identify the top 5% of the anomaly scores as anomalous. If you do not specifyThresholdParameter, the detector sets the threshold to 0.01."max","mean", or"median"— SpecifyThresholdParameteras a positive numeric scalar. If you do not specifyThresholdParameter, the detector sets the threshold to 1."customFunction"or"manual"—ThresholdParameterdoes not apply.
Function to compute custom detection threshold, specified as a function handle.
This argument applies only when ThresholdMethod is
specified as "customFunction".
The input to the function must be the vector of the anomaly scores.
The function must return a positive scalar corresponding to the detection threshold.
Isolation Forest Model Options
Number of trees in the isolation forest, specified as a positive integer.
The average path lengths used by the isolation forest algorithm to compute anomaly scores usually converge well before growing 100 isolation trees for both normal points and anomalies.
Number of observations to draw from the training data, without replacement, for each isolation tree, specified as a positive integer greater than or equal to 3.
The isolation forest algorithm performs well with a small value for
NumObservationsPerLearner, because using a small sample size
helps to detect dense anomalies and anomalies close to normal points. However, you
need to experiment with the sample size if N, the number of
training observations, is small.
Normalization
Normalization technique for training and testing, specified as
"zscore", "range", or
"off".
"range"— Rescale the data range to [0,1]."zscore"— Distance from a data point to the mean in terms of standard deviation"off"— Do not normalize the data.
For more information on normalization methods, see normalize.
Output Arguments
Anomaly detector model, returned as an TimeSeriesIForestDetector object.
Version History
Introduced in R2026a
See Also
train | detect | plot | plotHistogram | updateDetector
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Web サイトの選択
Web サイトを選択すると、翻訳されたコンテンツにアクセスし、地域のイベントやサービスを確認できます。現在の位置情報に基づき、次のサイトの選択を推奨します:
また、以下のリストから Web サイトを選択することもできます。
最適なサイトパフォーマンスの取得方法
中国のサイト (中国語または英語) を選択することで、最適なサイトパフォーマンスが得られます。その他の国の MathWorks のサイトは、お客様の地域からのアクセスが最適化されていません。
南北アメリカ
- América Latina (Español)
- Canada (English)
- United States (English)
ヨーロッパ
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)