nlssinit
Initialize nonlinear state-space model using measured time-domain system data
Since R2026a
Syntax
Description
nssInitialized = nlssinit(U,Y,nss)U and Y,
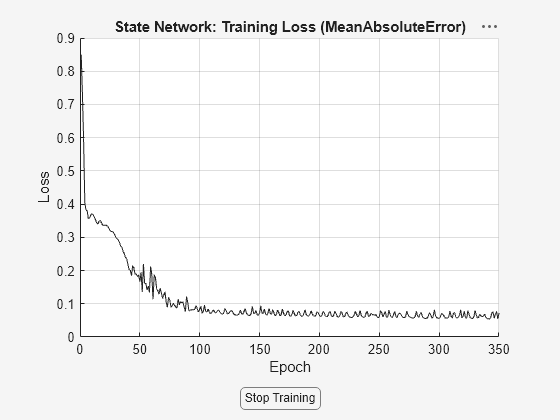
and default training options, to train the state and output networks of the idNeuralStateSpace
object nss. It estimates the weights and biases of the networks by
numerically approximating the state derivatives and performing an open-loop training. The
open-loop training minimizes the state-derivative prediction error for continuous-time
models and the state-update prediction error for discrete-time models. This syntax returns
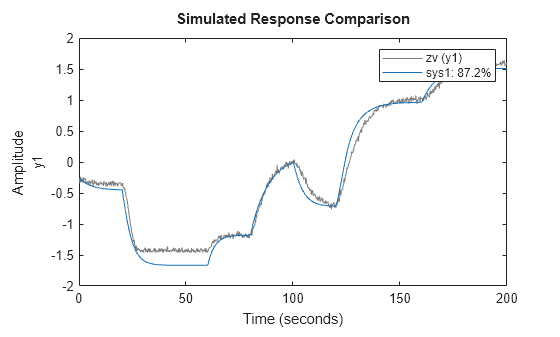
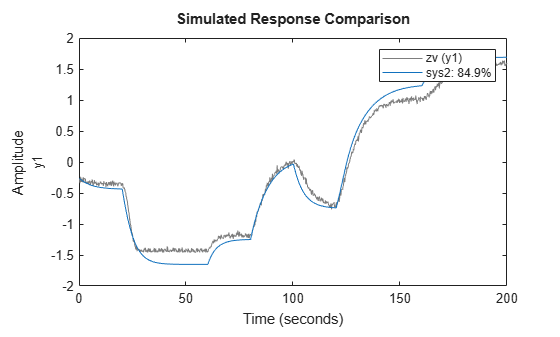
the idNeuralStateSpace object nssInitialized with the
trained state and output networks. You can use nssInitialized as the
initial model when estimating neural state-space models using nlssest.
nssInitialized = nlssinit(Data,nss)Data, and the default
training options, to train the state and output networks of nss.
nssInitialized = nlssinit(___,Options)
nssInitialized = nlssinit(___,Name=Value)
[
returns model parameters corresponding to the final loss and minimal training loss. If
nssInitialized,params] = nlssinit(___)UseLastExperimentForValidation is true, it also returns the
model parameters corresponding to minimal validation loss.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced in R2026a