結果:
I rarely use MATLAB.
10%
use MATLAB almost every day.
55%
use MATLAB once every 2-3 days.
10%
only use when specific task require
25%
20 票
For the last day or two, I've been getting "upstream" and other various errors on Answers. Seems to come and go. Anyone else having similar issues?
You are not a jedi yet !
20%
We not grant u the rank of master !
0%
Ready are u? What knows u of ready?
0%
May the Force be with you !
80%
5 票
In a discussion on LInkedin about my recent blog post, Do these 3 things to increase the reach of your open source MATLAB toolbox, I was asked by "Could you elaborate on why someone might consider opening/sharing their code? Thinking of early-career researchers, what might be in it for them?"
I'll give my answer here but I'm more interested in yours. How would you have answered this?
This is what I said:
- It's the right thing to do scientifically. A computational paper is essentially just an advertisement of what you've done. The code contains vital details about how you actually did it. A computational paper is incomplete without the code.
- If you only describe your algorithm in a paper, I have to implement it before I can apply your research to my problem. If you share the code, I can get started much more quickly using your research. This means I publish faster and since I am a good scientist, this means you get cited faster.
- Other scientists start off as users of your code. This leads to citations. Over time, some of them start deeply using and modifying your code, this leads to collaborators.
- Once you decide to share code via something like GitHub, you quickly start adopting good software engineering practices without initially realizing it. This improves the quality of your research since adopting good software practices makes it more likely that your software will give the right answers.
That last point can be a little hard to get your head around sometimes. Even if all you do is use file upload to get your stuff onto GitHub (i.e. you're not using git properly yet) you will start to naturally converge towards better code.
Why? Because as soon as you share code, you have to solve the problem of getting it to run on someone else's machine.
A trivial example concerns hard coded paths, for example. If you only ever run it on your machine then having a line like datafile = "C:\Mystuff\data.csv" always works but it breaks as soon as I try to run it on my machine. You'll look at this and think "Maybe there's a better way to do that".
Similarly dependencies. Your Path may be full of stuff that isn't present on my machine. As soon as I try to run your code, it won't work and you'll have to figure out how to handle dependencies in a reproducible way.
Documentation! An empty README.md is no good if you expect me to know how to use your code. You at least have to say something like "To run this, type runme(N) into MATLAB where N is the size of the model...etc etc)
The act of sharing, and dealing with the consequences, leads to much better code than if you keep it to yourself.
The MATLAB R2025a release gave figures a makeover. @Brian Knolhoff, a developer on the Figure Infrastructure and Services team, reviews the new Figure Container in the Graphics and App Building blog.
Learn the four ways to tile figures, what docking means, and other new features.
In case you missed it in my overview of the MATLAB R2025a release, Markdown support has been greatly improved. This picture says it all

I saw this on Reddit and thought of the past mini-hack contests. We have a few folks here who can do something similar with MATLAB.

I had an error in the web version Matlab, so I exited and came back in, and this boy was plotted.
It seems like the financial news is always saying the stock market is especially volatile now. But is it really? This code will show you the daily variation from the prior day. You can see that the average daily change from one day to the next is 0.69%. So any change in the stock market from the prior day less than about 0.7% or 1% is just normal "noise"/typical variation. You can modify the code to adjust the starting date for the analysis. Data file (Excel workbook) is attached (hopefully - I attached it twice but it's not showing up yet).

% Program to plot the Dow Jones Industrial Average from 1928 to May 2025, and compute the standard deviation.
% Data available for download at https://finance.yahoo.com/quote/%5EDJI/history?p=%5EDJI
% Just set the Time Period, then find and click the download link, but you ned a paid version of Yahoo.
%
% If you have a subscription for Microsoft Office 365, you can also get historical stock prices.
% Reference: https://support.microsoft.com/en-us/office/stockhistory-function-1ac8b5b3-5f62-4d94-8ab8-7504ec7239a8#:~:text=The%20STOCKHISTORY%20function%20retrieves%20historical,Microsoft%20365%20Business%20Premium%20subscription.
% For example put this in an Excel Cell
% =STOCKHISTORY("^DJI", "1/1/2000", "5/10/2025", 0, 1, 0, 1,2,3,4, 5)
% and it will fill out a table in Excel
%====================================================================================================================
clc; % Clear the command window.
close all; % Close all figures (except those of imtool.)
imtool close all; % Close all imtool figures if you have the Image Processing Toolbox.
clear; % Erase all existing variables. Or clearvars if you want.
workspace; % Make sure the workspace panel is showing.
format long g;
format compact;
fontSize = 14;
filename = 'Dow Jones Industrial Index.xlsx';
data = readtable(filename);
% Date,Close,Open,High,Low,Volume
dates = data.Date;
closing = data.Close;
volume = data.Volume;
% Define start date and stop date
startDate = datetime(2011,1,1)
stopDate = dates(end)
selectedDates = dates > startDate;
% Extract those dates:
dates = dates(selectedDates);
closing = closing(selectedDates);
volume = volume(selectedDates);
% Plot Volume
hFigVolume = figure('Name', 'Daily Volume');
plot(dates, volume, 'b-');
grid on;
xticks(startDate:calendarDuration(5,0,0):stopDate)
title('Dow Jones Industrial Average Volume', 'FontSize', fontSize);
hFig = figure('Name', 'Daily Standard Deviation');
subplot(3, 1, 1);
plot(dates, closing, 'b-');
xticks(startDate:calendarDuration(5,0,0):stopDate)
drawnow;
grid on;
caption = sprintf('Dow Jones Industrial Average from %s through %s', dates(1), dates(end));
title(caption, 'FontSize', fontSize);
% Get the average change from one trading day to the next.
diffs = 100 * abs(closing(2:end) - closing(1:end-1)) ./ closing(1:end-1);
subplot(3, 1, 2);
averageDailyChange = mean(diffs)
% Looks pretty noisy so let's smooth it for a nicer display.
numWeeks = 4;
diffs = sgolayfilt(diffs, 2, 5*numWeeks+1);
plot(dates(2:end), diffs, 'b-');
grid on;
xticks(startDate:calendarDuration(5,0,0):stopDate)
hold on;
line(xlim, [averageDailyChange, averageDailyChange], 'Color', 'r', 'LineWidth', 2);
ylabel('Percentage', 'FontSize', fontSize);
caption = sprintf('Day-to-Day Change Percentage. Average Daily Change (from prior day) = %.2f%%', averageDailyChange);
title(caption, 'FontSize', fontSize);
drawnow;
% Get the stddev over a 5 trading day window.
sd = stdfilt(closing, ones(5, 1));
% Get it relative to the magnitude.
sd = sd ./ closing * 100;
averageVariation = mean(sd)
numWeeks = 2;
% Looks pretty noisy so let's smooth it for a nicer display.
sd = sgolayfilt(sd, 2, 5*numWeeks+1);
% Plot it.
subplot(3, 1, 3);
plot(dates, sd, 'b-');
grid on;
xticks(startDate:calendarDuration(5,0,0):stopDate)
hold on;
line(xlim, [averageVariation, averageVariation], 'Color', 'r', 'LineWidth', 2);
ylabel('Percentage', 'FontSize', fontSize);
caption = sprintf('Weekly Standard Deviation, Averaged Over %d Weeks (%d trading days). Mean SD = %.2f', ...
numWeeks, 5*numWeeks+1, averageVariation);
title(caption, 'FontSize', fontSize);
% Maximize figure window.
g = gcf;
g.WindowState = 'maximized';
The following lines were added to the subplot function in version 2025a (line 291):
if ancestorFigure.Units == "normalized"
waitfor(ancestorFigure,'FigureViewReady',true);
end
That code isn't in version 2024a.
Because of this, I'm experiencing issues that cause the code to stop running when using subplot in this way:
figure('Units','normalized','Position',[0 0 0.3 0.3])
subplot(1,2,1)
...
Has anyone else encountered this error?
Does anyone understand the need for those lines of code?
As far as I can tell, there is still no official support for creating publication-ready tables from regression output, either as latex or natively. Although MATLAB isn't primarily statistical software, this still seems like an oversight, as almost any similar software has this capability built-in or as a package.
Due to MATLAB being banned in some mainland Chinese universities in 2020, in recent years a Chinese company called "Suzhou Tongyuan SoftControl" has completely imitated MATLAB’s behavior. Below are some screenshots as evidence. What is your opinion on this issue?
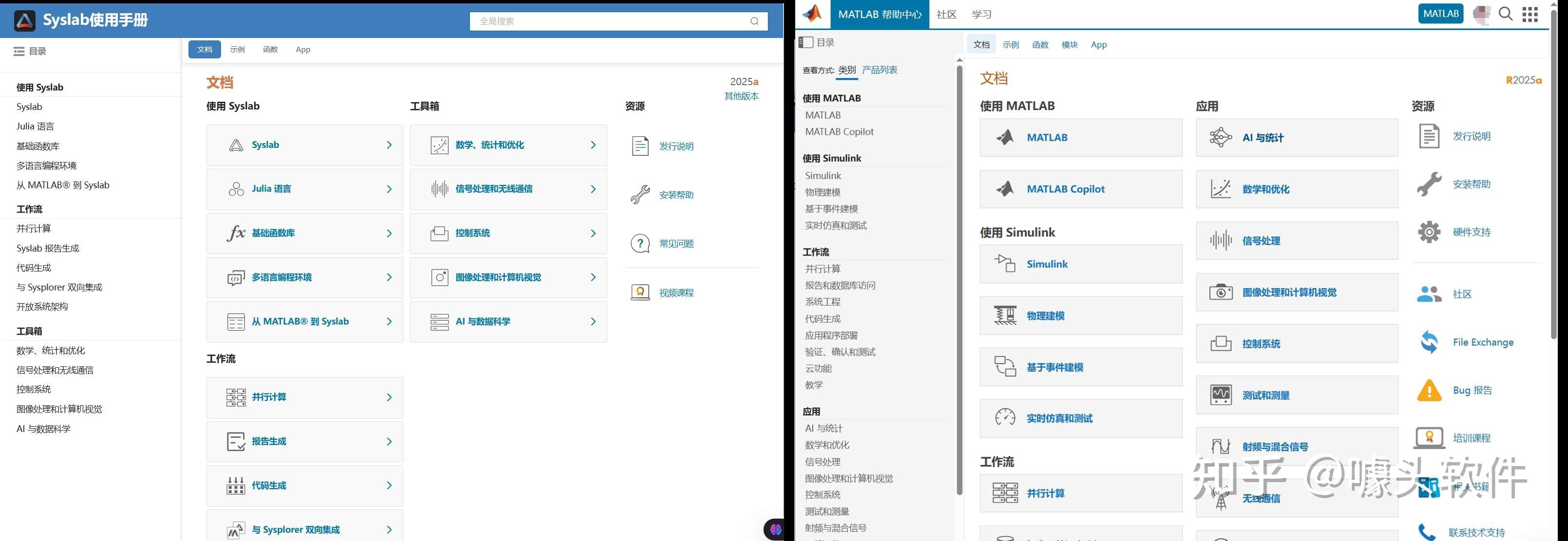
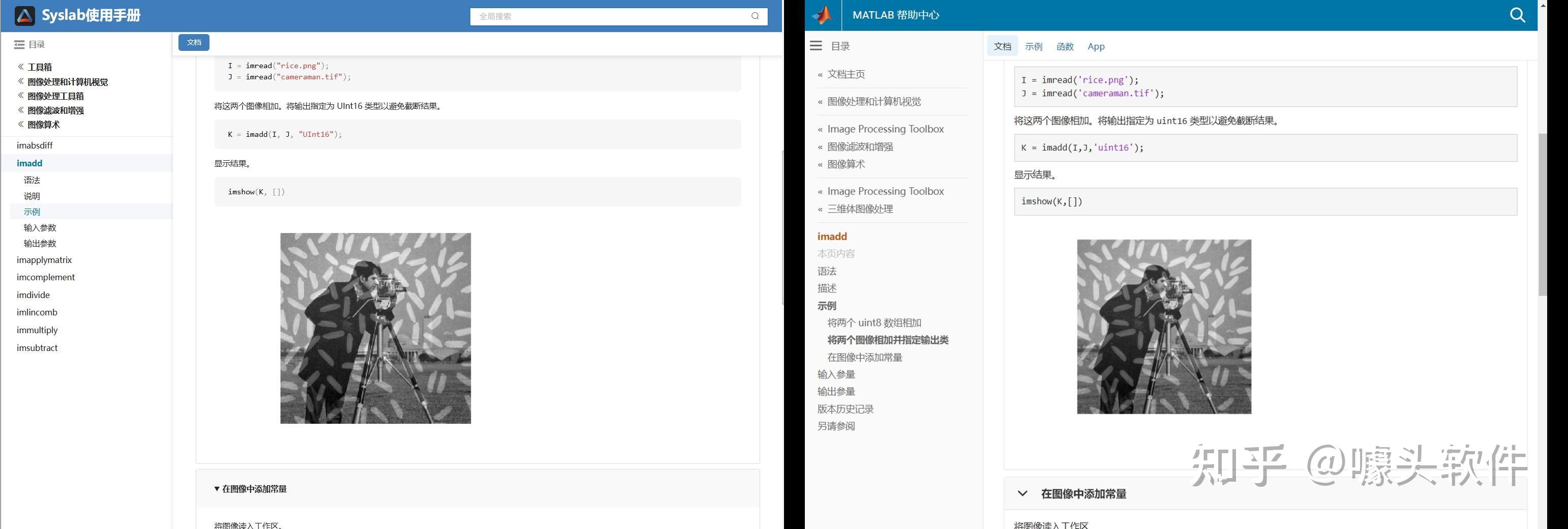
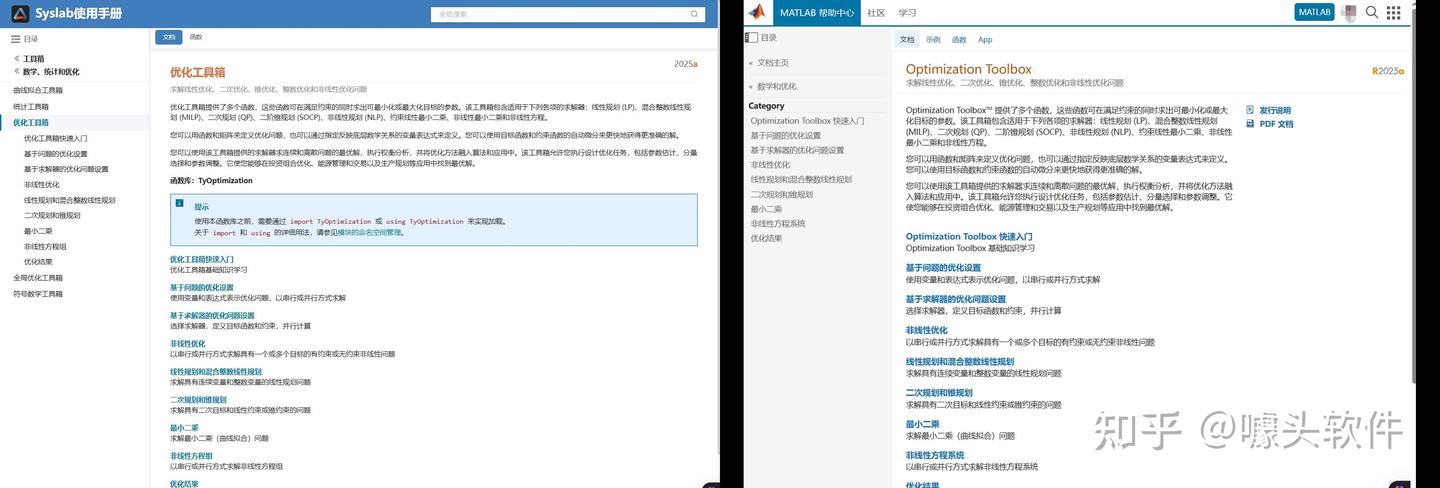
Source: Syslab 帮助文档-苏州同元软控
After waiting for a long time, the MathWorks official Community has finally resumed some of its functionalitys! Congratulations! Next, I’d like to share some thoughts to help prevent such outages from happening again, as they have affected far too many people.
- Almost all resources rely solely on MathWorks servers. Once a failure (or a ransomware attack) occurs, everything is paralyzed, and there isn’t even a temporary backup server? For a big company like MathWorks to have no contingency plan at all is eye-opening. This tells us that we should have our own temporary emergency servers!
- The impact should be minimized. For example, many users need to connect to the official servers to download various support packages, such as the “Deep Learning Toolbox Converter for ONNX Model Format.” Could these be backed up and mirrored to the “releases” section of a GitHub repository, so users in need can download them.
- A large proportion of users who have already installed MATLAB cannot access the online help documentation. Since R2023a, installing the help documentation locally has become optional. This only increases the burden on the servers? Moreover, the official website only hosts documentation for the past five years. That means after 2028, if I haven’t installed the local offline documentation, I won’t be able to access the online documentation for R2023a anymore?
Anything else you’d like to add? Feel free to leave a comment.
Any status updates on the license center and add on tool boxes?
@William Rose, Your dedication to helping others and sharing your knowledge is a big win for the community. Thanks for taking the time to contribute so thoughtfully - your impact is definitely being noticed.👏
Keep it up!
The attached code is an animated solution of the three body problem. On 2024b it runs perfectly fine. When we tried it on 2025a, the animation constantly hitches, the CPU usage is almost double and the runtime is much slower. The curves also look less detailed and jagged in some places. When we run it without drawing anything, the performance seems comparable between versions, but still slightly slower. All of this behavior persists across different hardware. Anybody else having this kind of problem with the new release? I'm suspecting the graphics backend changes may be the culprit here...
clc
clear
close
syms t x1(t) y1(t) x2(t) y2(t) x3(t) y3(t)
G = 6.6743 * 10^-11;
%epsilon = 1e-4
m1 = 10^12;
m2 = 1*10^12;
m3 = 1*10^12;
r1 = [x1(t),y1(t)];
K1 = 1/2 * m1 * (diff(x1(t),t)^2 + diff(y1(t),t)^2);
r2 = [x2(t),y2(t)];
K2 = 1/2 * m2 * (diff(x2(t),t)^2 + diff(y2(t),t)^2);
r3 = [x3(t),y3(t)];
K3 = 1/2 * m3 * (diff(x3(t),t)^2 + diff(y3(t),t)^2);
L1x = diff(diff(K1,diff(x1(t),t)) , t);
L1y = diff(diff(K1,diff(y1(t),t)) , t);
L2x = diff(diff(K2,diff(x2(t),t)) , t);
L2y = diff(diff(K2,diff(y2(t),t)) , t);
L3x = diff(diff(K3,diff(x3(t),t)) , t);
L3y = diff(diff(K3,diff(y3(t),t)) , t);
r12 = r2 - r1;
r13 = r3 - r1;
r23 = r3 - r2;
dlugosc_r12 = sqrt(r12(1)^2 + r12(2)^2);
dlugosc_r13 = sqrt(r13(1)^2 + r13(2)^2);
dlugosc_r23 = sqrt(r23(1)^2 + r23(2)^2);
Q12 = G * m1 * m2 / dlugosc_r12^2 * (r2-r1)/dlugosc_r12;
Q13 = G * m1 * m3 / dlugosc_r13^2 * (r3-r1)/dlugosc_r13;
Q23 = G * m2 * m3 / dlugosc_r23^2 * (r3-r2)/dlugosc_r23;
Q21 = -Q12;
Q32 = -Q23;
Q31 = -Q13;
Q1 = Q12 + Q13;
Q2 = Q21 + Q23;
Q3 = Q31 + Q32;
eqn_1_x = L1x == Q1(1);
eqn_1_y = L1y == Q1(2);
eqn_2_x = L2x == Q2(1);
eqn_2_y = L2y == Q2(2);
eqn_3_x = L3x == Q3(1);
eqn_3_y = L3y == Q3(2);
syms X1 Y1 X2 Y2 X3 Y3
Q1_num = subs(Q1,[x1(t), y1(t), x2(t), y2(t), x3(t), y3(t)],[X1, Y1, X2, Y2, X3, Y3]);
Q2_num = subs(Q2,[x1(t), y1(t), x2(t), y2(t), x3(t), y3(t)],[X1, Y1, X2, Y2, X3, Y3]);
Q3_num = subs(Q3,[x1(t), y1(t), x2(t), y2(t), x3(t), y3(t)],[X1, Y1, X2, Y2, X3, Y3]);
syms vx1 vy1 vx2 vy2 vx3 vy3
state_dot = [
vx1;
vy1;
vx2;
vy2;
vx3;
vy3;
Q1_num(1)/m1;
Q1_num(2)/m1;
Q2_num(1)/m2;
Q2_num(2)/m2;
Q3_num(1)/m3;
Q3_num(2)/m3
];
f = matlabFunction(state_dot, 'Vars', {sym('t'), [X1; Y1; X2; Y2; X3; Y3; vx1; vy1; vx2; vy2; vx3; vy3]});
u0 = [-1e5; %x1
0; %y1
1e5; %x2
0; %y2
0; %x3
sqrt(3)*1e5; %y3
-11/2 * 1e-3; %vx1
11/2*sqrt(3)*1e-3; %vy1
-11/2 * 1e-3; %vx2
-11/2*sqrt(3)*1e-3; %vy2
11e-3; %vx3
0]; %vy3
tspan = [0, 1e9];
%options = odeset('RelTol', 1e-15, 'AbsTol', 1e-20);
[t_sol, u_sol] = ode45(f, tspan, u0);
t_anim = linspace(t_sol(1), t_sol(end), 5000);
u_anim = interp1(t_sol, u_sol, t_anim);
%%
% figure;
tor_1 = plot(u_anim(:,1), u_anim(:,2), 'r', 'LineWidth',1.5); hold on;
tor_2 = plot(u_anim(:,3), u_anim(:,4), 'g', 'LineWidth',1.5);
tor_3 = plot(u_anim(:,5), u_anim(:,6), 'b', 'LineWidth',1.5);
% xlabel('x [m]');
% ylabel('y [m]');
% legend('Ciało 1', 'Ciało 2', 'Ciało 3');
% title('Trajektorie ciał w układzie trzech ciał');
% axis equal
% grid on;
pozycja_1 = plot(u_anim(1,1),u_anim(1,2),'ro','markersize',10,'markerface','r'); hold on
pozycja_2 = plot(u_anim(1,3),u_anim(1,4),'go','markersize',10,'markerface','g');
pozycja_3 = plot(u_anim(1,5),u_anim(1,6),'bo','markersize',10,'markerface','b');
% xlim([-2e5,2e5])
% ylim([-2e5,2e5])
axis equal
for i = 1 : 1 : length(t_sol)
set(pozycja_1,'XData', u_anim(i,1),'YData', u_anim(i,2));
set(pozycja_2,'XData', u_anim(i,3),'YData', u_anim(i,4));
set(pozycja_3,'XData', u_anim(i,5),'YData', u_anim(i,6));
set(tor_1,'XData', u_anim(1:i,1),'YData', u_anim(1:i,2));
set(tor_2,'XData', u_anim(1:i,3),'YData', u_anim(1:i,4));
set(tor_3,'XData', u_anim(1:i,5),'YData', u_anim(1:i,6));
drawnow;
% pause(0.001);
end
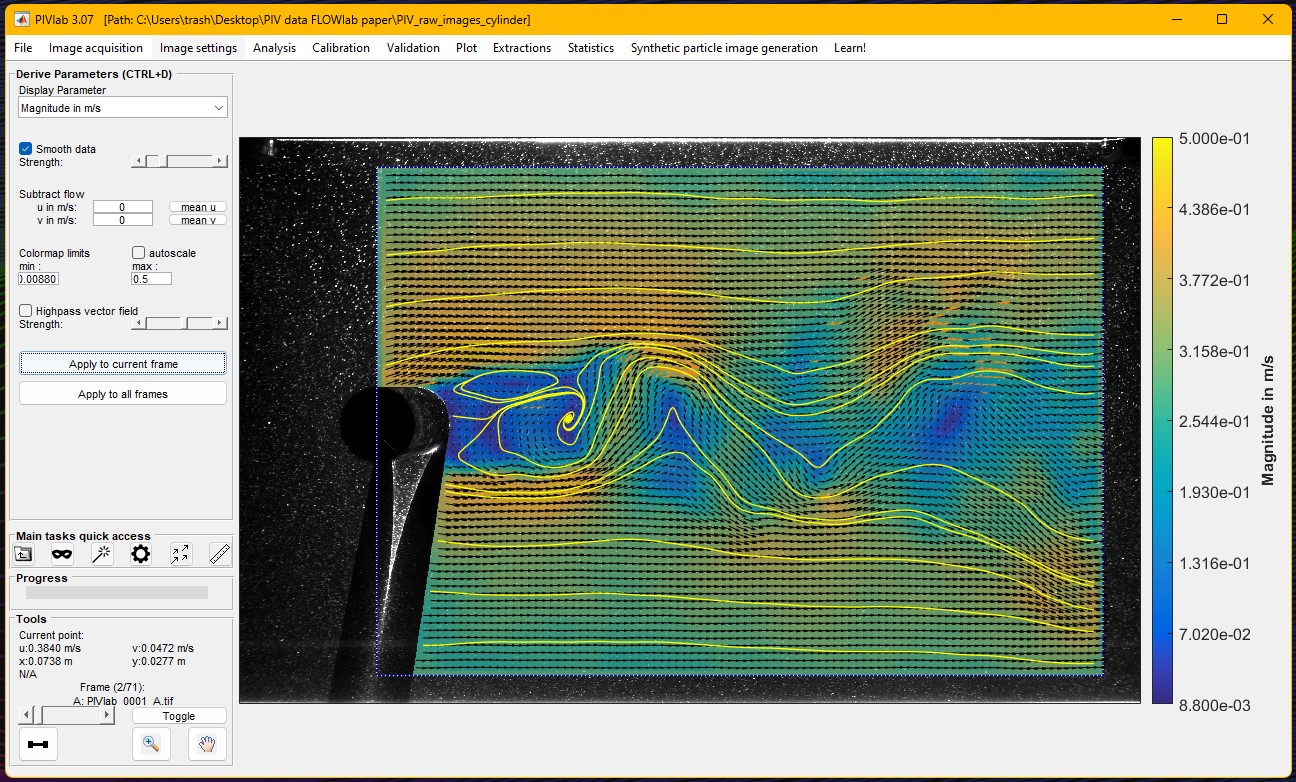
During the past twelve months, PIVlab, a MATLAB Community Toolbox for particle interference velocimetry (a technique for fluid flow measurement) set a new record for all-time File Exchange downloads, surpassing one hundred thousand, dating back to 2010. It also recently eclipsed the 1000 downloads/month mark on File Exchange.
Congratulations to @William Thielicke and his team for this fantastic long term achievement and head over to the File Exchange to download and use it: PIVlab - particle image velocimetry (PIV) tool with GUI - File Exchange - MATLAB Central
Why is RoBERTa not available as a pretrained model? It is superior to BERT in many fields and has become more popular in the literature. For faster inference, you should offer DistilBERT, which is more modern than BERT but smaller/faster. The respository hasn't been updated in two years, which is a lifetime in the field of deep learning.
https://github.com/matlab-deep-learning/transformer-models
I wanted to turn a Markdown nested list of text labels:
- A
- B
- C
- D
- G
- H
- E
- F
- Q
into a directed graph, like this:

Here is my blog post with some related tips for doing this, including text I/O, text processing with patterns, and directed graph operations and visualization.