pcares
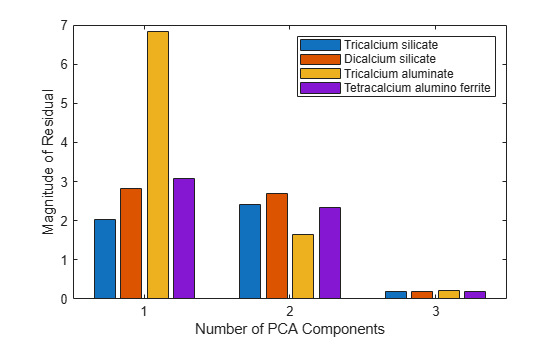
Residuals from principal component analysis
Description
residuals = pcares(X,NumComponents)NumComponents principal
components of the data matrix X.
pcares does not normalize the columns of X. You
can perform principal component analysis based on standardized variables using
pcares(zscore(X),NumComponents). To perform principal component
analysis directly on a covariance or correlation matrix, but without computing residuals,
use pcacov.
[
additionally returns an approximation to residuals,reconstructed] = pcares(X)X obtained by retaining its
first NumComponents principal components.
reconstructed is equal to X minus
residuals.
Examples
Input Arguments
Output Arguments
References
[1] Jackson, J. E. A User's Guide to Principal Components, John Wiley and Sons, 1991.
[2] Jolliffe, I. T. Principal Component Analysis, 2nd Edition, Springer, 2002.
[3] Krzanowski, W. J. Principles of Multivariate Analysis: A User's Perspective. New York: Oxford University Press, 1988.
[4] Seber, G. A. F. Multivariate Observations. Hoboken, NJ: John Wiley & Sons, Inc., 1984.
Version History
Introduced before R2006a