結果:
There is a communication regarding "How can I set the text font style of a Data Cursor object interactively on a plot?". But I am not clear on the answer found in this link:
https://www.mathworks.com/matlabcentral/answers/95968-how-can-i-set-the-text-font-style-of-a-data-cursor-object
I do not know how and where to put the recommended commands. Would you please clarfity and give me more details?
Thank you.
Worth the wait: seven new online training courses and one new learning path were released with 25a, covering topics in Controls, Electrification, and Physical Modeling. This release also brings new functionality to support interactions across both MATLAB and Simulink within a single course, beginning with the new Controls courses below:
- Control System Design with MATLAB and Simulink (learning path – includes the 5 controls courses listed below)
- Control System Modeling Essentials
- Linearization of Nonlinear Systems
- Control System Analysis Techniques
- PID Control Techniques
- Classical Controller Design Techniques
- Battery State Estimation
- Motor Modeling with Simscape Electrical
Did you know that function double with string vector input significantly outperforms str2double with the same input:
x = rand(1,50000);
t = string(x);
tic; str2double(t); toc
tic; I1 = str2double(t); toc
tic; I2 = double(t); toc
isequal(I1,I2)
Recently I needed to parse numbers from text. I automatically tried to use str2double. However, profiling revealed that str2double was the main bottleneck in my code. Than I realized that there is a new note (since R2024a) in the documentation of str2double:
"Calling string and then double is recommended over str2double because it provides greater flexibility and allows vectorization. For additional information, see Alternative Functionality."
mlapp being a binary is a pain point for source control. It means that you either have to:
- have hooks in your source control system to zip/unzip a mlapp. However, The Mathworks have informed users not to rely on this as the mlapp format may change.
- do all your source control in MATLAB. This is non standard behaviour. Source code and source control should be independent of each other. Web front-ends to source control systems, 3rd party source control apps, CI/CD systems and much more are extremely limited in what they can do with mlapps.
I wish an mlapp could just be a directory full of the required text/other files.
Requested to post this here from reddit.
There is no call to rescan audio devices in audioPlayerRecorder, even though PortAudio has such a call. I have a measurement environment that takes a long time to initialise. If I forget to plug in my audio device, I have to do it all over again...
This just came out. @Michelle Hirsch spoke to Jousef Murad and answer his questions about the big change in the desktop in R2025a and explained what was going on behind the scene. Enjoy!
The Big MATLAB Update: Dark Mode, Cloud & the Future of Engineering - Michelle Hirsch
Share your ideas, suggestions, and wishlists for improving MathWorks products. What would make the software absolutely perfect for you? Discuss your idea(s) with other community users.
Guidelines & Tips
We encourage all ideas, big or small! To help everyone understand and discuss your suggestion, please include as much detail as possible in your post:
- Product or Feature: Clearly state which product (e.g., MATLAB, Simulink, a toolbox, etc.) or specific feature your idea relates to.
- The Problem or Opportunity: Briefly describe what challenge you’re facing or what opportunity you see for improvement.
- Your Idea: Explain your suggestion in detail. What would you like to see added, changed, or improved? How would it help you and other users?
- Examples or Use Cases (optional): If possible, include an example, scenario, or workflow to illustrate your idea.
- Related Posts (optional): If you’ve seen similar ideas or discussions, feel free to link to them for context.
Ready to share your idea?
Click here and then "Start a Discussion”, and let the community know how MATLAB could be even better for you!
Thank you for your contributions and for helping make MATLAB Central a vibrant place for sharing and improving ideas.
These got released last week and the process for using them on your local machine with MATLAB is very similar to how you use the local deepseek models as I demonstrated in my February blog post How to run local DeepSeek models and use them with MATLAB » The MATLAB Blog - MATLAB & Simulink
You need Ollama and the LLMs with MATLAB package installed (Details on how to do this in the blog post above). Then you run the following in your operating systems' command line
ollama pull gpt-oss:20b
Over to MATLAB and set up a chat session
>> chat = ollamaChat("gpt-oss:20b")
chat =
ollamaChat with properties:
ModelName: "gpt-oss:20b"
Endpoint: "127.0.0.1:11434"
TopK: Inf
MinP: 0
TailFreeSamplingZ: 1
Temperature: 1
TopP: 1
StopSequences: [0×0 string]
TimeOut: 120
SystemPrompt: []
ResponseFormat: "text"
FunctionNames: []
txt = generate(chat,"Who are you?")
txt =
"I’m ChatGPT – a conversational AI developed by OpenAI. My core is the GPT‑4 language model, which has been trained on a massive mix of text from books, websites, articles and other sources to understand and generate human‑like language. I don’t have feelings, consciousness, or a personal identity; I’m a tool that can help answer questions, brainstorm ideas, explain concepts, draft text, and more. My goal is to understand the context you give me and respond in a helpful, accurate and safe way. If there’s something specific you’d like to know or do, just let me know!"
This is the smaller of the two, new open models and it is bringing my aging desktop to its knees. My GPU is too small to do the work so I think everything is happening on the CPU and its slooooow. Will try on my Mac next
Let me know if you try this out!
Long before I joined MathWorks, I was a member of the academic Research Software Engineering (RSE) community where part of my mission was to introduce basic software engineering concepts to the research community. Things like version control, testing and even simply writing code instead of using only pointy-clicky GUIs before copying and pasting the results plot into a word document. I've seen things..........*shudders*
The RSE movement is still going very strong and I am elated that MathWorks is increasingly interacting with it. One example of such interaction is a video tutorial contributed by my colleauge @Mihaela Jarema to a comminity seminar series called 'A summer of Testing' It's linked to below
The video assumes you've never run a test before and gently guides you through the principles. Along the way you'll learn about some of MATLAB's superb testing capabilities. Things like
- Unit testing Framework
- Test Browser App
- Code Coverage
- Test Fixtures (Setup and teardown)
- Test driven devellopment
- Function argument validation
- CI/CD using GitHub actions
Go check out out.
I just wanted here to share a link to some .gif animations I created over the years with Matlab :-)
I think gif animations are great supports for scientific diffusion.
Hello to all!
I would like to share with the Matlab and Simulink community this video about Neural Networks in Simulink.
This is a series of videos that use a multilayer perceptron implemented in Simulink as a case study. Why Simulink? Because it's a visual and intuitive modeling tool, you can see the forward propagation of this network and better understand the flow. The objective of this series is to show the implementation using Simulink for both simulation and Arduino, as well as its training using Matlab and Matlab with Deep Learning Toolbox, and a video of training with Python.
The video is in Spanish, but the Simulink model is available in English for the entire community; subtitles are also available.
The files are located in the first comment of each video. We hope you find it interesting and enjoyable. Best regards!
Here I share the link to the first video.
In many parts of Africa, particularly in technical universities and engineering institutes, physical laboratories are scarce or poorly equipped. This reality deeply limits the hands-on experience students deserve, especially in fields like control systems, signal processing, power electronics, and fluid mechanics.
But MATLAB and Simulink can fill part of this gap.
As an educator and researcher, I’ve made it my mission to promote MATLAB as a didactic simulation environment that brings real-world experimentation into the virtual space—affordable, accessible, and scalable. Whether simulating dynamic systems, visualizing electromagnetic fields, or tuning PID controllers interactively, students can develop strong intuition without needing costly hardware.
🔧 I’ve used MATLAB to teach:
- Power systems and control theory without needing real generators or oscilloscopes,
- Hydrology and environmental modeling without field sensors,
- Robotics and AI concepts even where no robot is available.
🌍 This is more than a tool for me. It’s a bridge between educational ambition and limited infrastructure.
I dream of creating MATLAB-based virtual laboratories across African institutions. And I know I’m not alone.
Is anyone else here working on similar goals in under-resourced regions? Let’s connect and make it real.
— Patrick K.N.
As someone who grew up programming in C#, I often find myself wishing for a tighter, more native integration between MATLAB and C#.
There’s so much I dream of doing—leveraging the power of Simulink models or MATLAB’s advanced numerical libraries inside my .NET desktop or web applications. Of course, I know there are some workarounds: COM automation, MATLAB Engine API for .NET, or using MATLAB Compiler SDK… but let’s be honest: it’s not quite as seamless as I’d hope.
I imagine a world where:
- I could directly call MATLAB functions from C# as if they were .NET assemblies, without middleware.
- Simulink blocks could generate portable C# code (not just C/C++).
- MATLAB UI components could be embedded in WPF/WinForms apps natively.
Until then... we make do with what we have. But the vision remains.
Anyone else here trying to bridge MATLAB and C# in their workflow? I’d love to hear your experiences or ideas!
— Patrick K.N.
I found some beautiful computational art made by a developer called @yuruyurau who used a language called Processing. Unfortunately, I know very little about this language so I asked Claude to convert it to MATLAB for me.
Give it a try yourself and show me what you come up with.
Details here: Pair programming with Claude to produce computational art in MATLAB » The MATLAB Blog - MATLAB & Simulink
I have started a blog series on the history of image display in MATLAB. If this topic interests you, and if there is something in particular you would like me to address in the series, let me know.
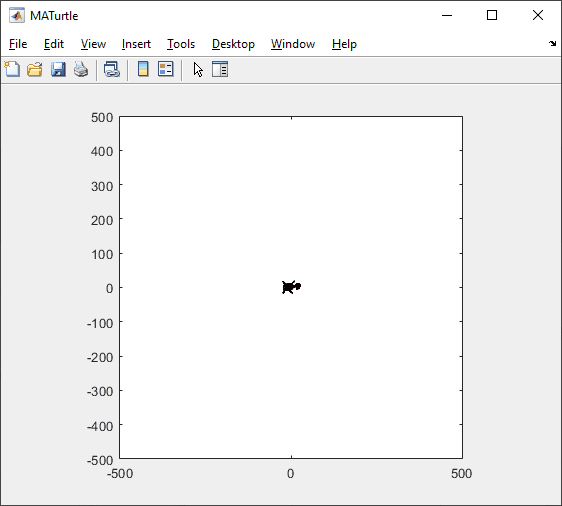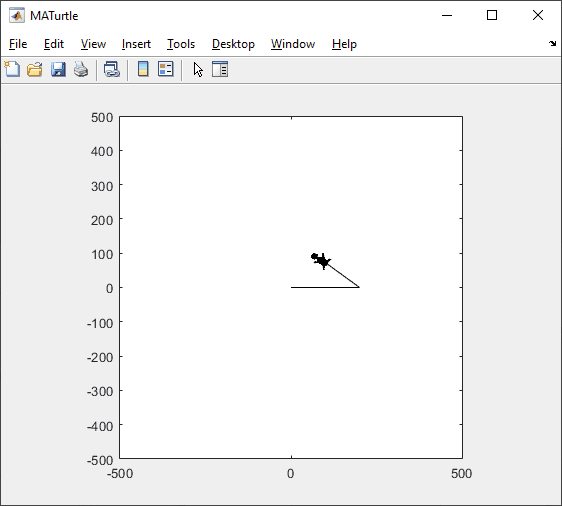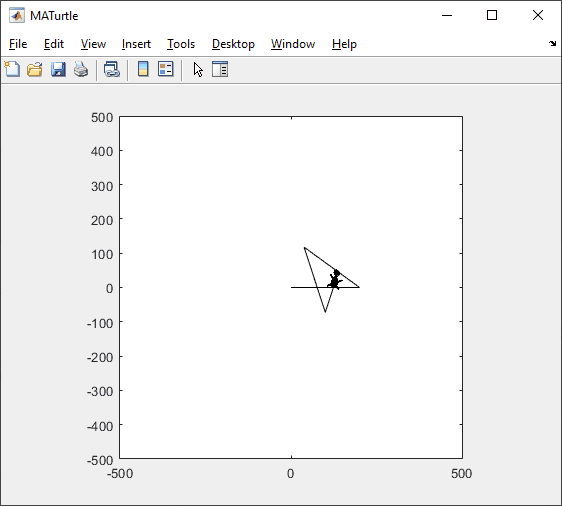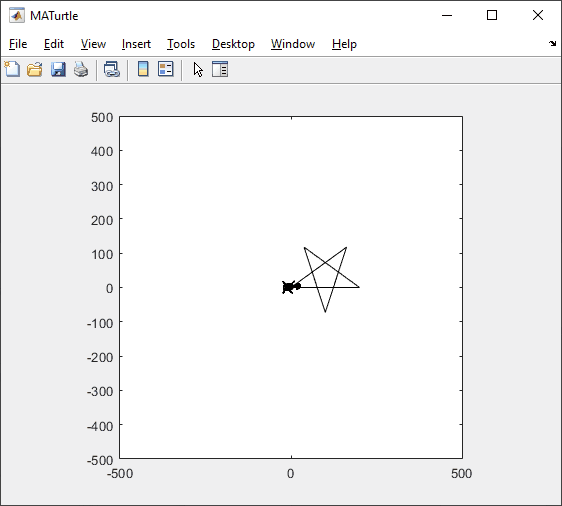
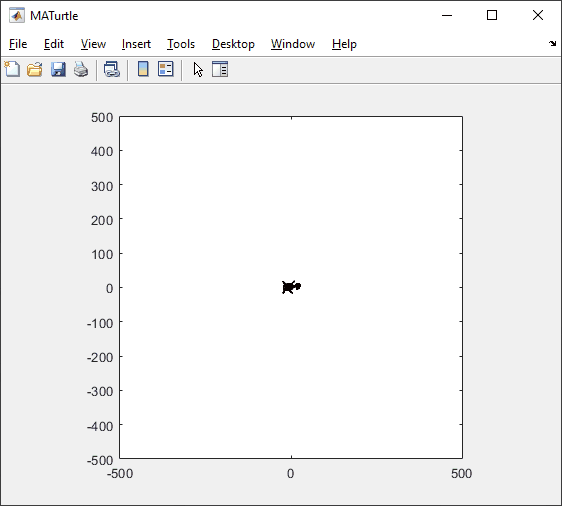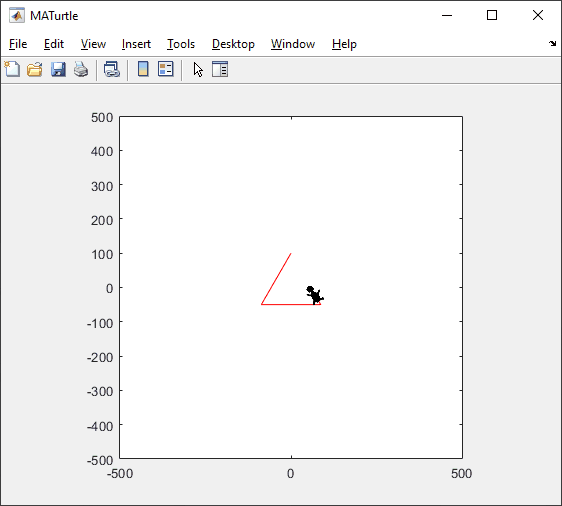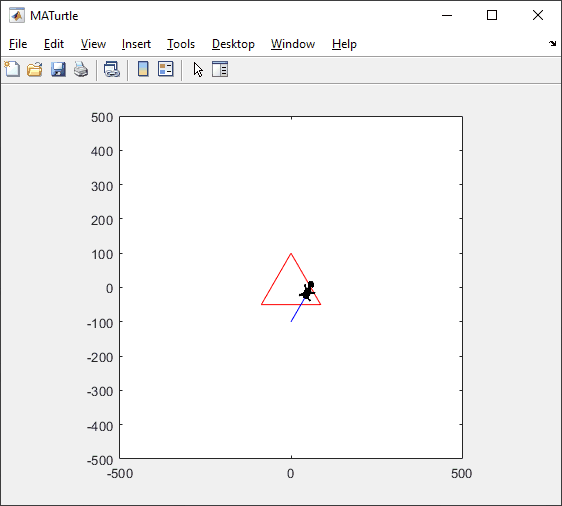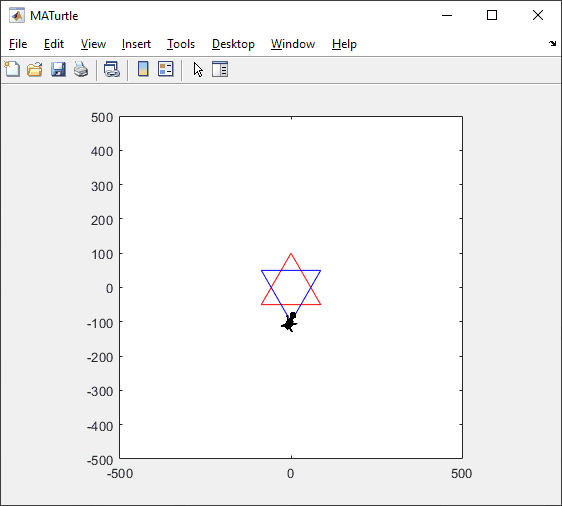
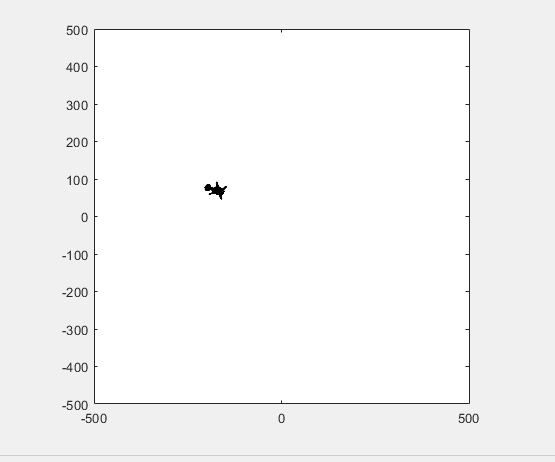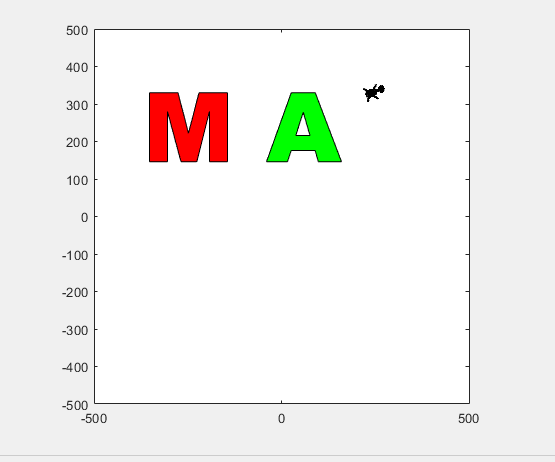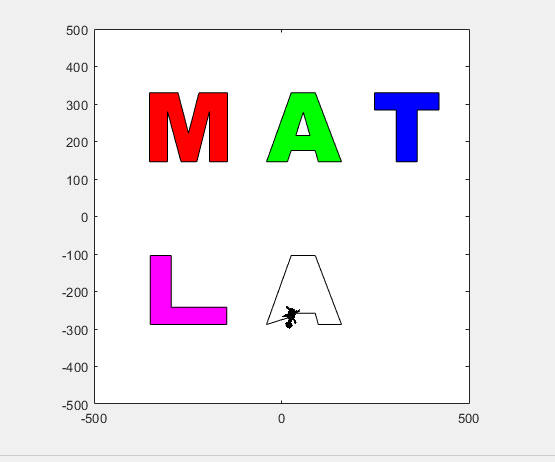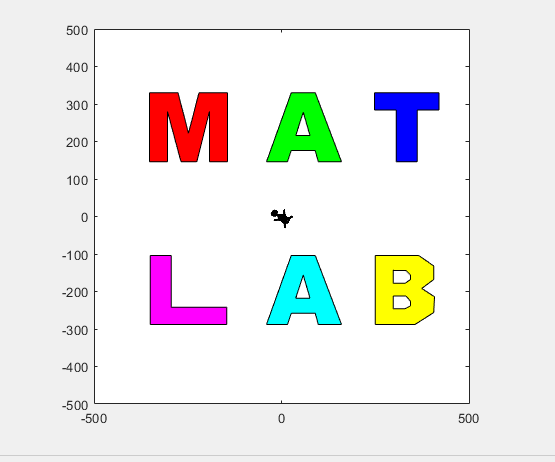
t = turtle(); % Start a turtle
t.forward(100); % Move forward by 100
t.backward(100); % Move backward by 100
t.left(90); % Turn left by 90 degrees
t.right(90); % Tur right by 90 degrees
t.goto(100, 100); % Move to (100, 100)
t.turnto(90); % Turn to 90 degrees, i.e. north
t.speed(1000); % Set turtle speed as 1000 (default: 500)
t.pen_up(); % Pen up. Turtle leaves no trace.
t.pen_down(); % Pen down. Turtle leaves a trace again.
t.color('b'); % Change line color to 'b'
t.begin_fill(FaceColor, EdgeColor, FaceAlpha); % Start filling
t.end_fill(); % End filling
t.change_icon('person.png'); % Change the icon to 'person.png'
t.clear(); % Clear the Axes
classdef turtle < handle
properties (GetAccess = public, SetAccess = private)
x = 0
y = 0
q = 0
end
properties (SetAccess = public)
speed (1, 1) double = 500
end
properties (GetAccess = private)
speed_reg = 100
n_steps = 20
ax
l
ht
im
is_pen_up = false
is_filling = false
fill_color
fill_alpha
end
methods
function obj = turtle()
figure(Name='MATurtle', NumberTitle='off')
obj.ax = axes(box="on");
hold on,
obj.ht = hgtransform();
icon = flipud(imread('turtle.png'));
obj.im = imagesc(obj.ht, icon, ...
XData=[-30, 30], YData=[-30, 30], ...
AlphaData=(255 - double(rgb2gray(icon)))/255);
obj.l = plot(obj.x, obj.y, 'k');
obj.ax.XLim = [-500, 500];
obj.ax.YLim = [-500, 500];
obj.ax.DataAspectRatio = [1, 1, 1];
obj.ax.Toolbar.Visible = 'off';
disableDefaultInteractivity(obj.ax);
end
function home(obj)
obj.x = 0;
obj.y = 0;
obj.ht.Matrix = eye(4);
end
function forward(obj, dist)
obj.step(dist);
end
function backward(obj, dist)
obj.step(-dist)
end
function step(obj, delta)
if numel(delta) == 1
delta = delta*[cosd(obj.q), sind(obj.q)];
end
if obj.is_filling
obj.fill(delta);
else
obj.move(delta);
end
end
function goto(obj, x, y)
dx = x - obj.x;
dy = y - obj.y;
obj.turnto(rad2deg(atan2(dy, dx)));
obj.step([dx, dy]);
end
function left(obj, q)
obj.turn(q);
end
function right(obj, q)
obj.turn(-q);
end
function turnto(obj, q)
obj.turn(obj.wrap_angle(q - obj.q, -180));
end
function pen_up(obj)
if obj.is_filling
warning('not available while filling')
return
end
obj.is_pen_up = true;
end
function pen_down(obj, go)
if obj.is_pen_up
if nargin == 1
obj.l(end+1) = plot(obj.x, obj.y, Color=obj.l(end).Color);
else
obj.l(end+1) = go;
end
uistack(obj.ht, 'top')
end
obj.is_pen_up = false;
end
function color(obj, line_color)
if obj.is_filling
warning('not available while filling')
return
end
obj.pen_up();
obj.pen_down(plot(obj.x, obj.y, Color=line_color));
end
function begin_fill(obj, FaceColor, EdgeColor, FaceAlpha)
arguments
obj
FaceColor = [.6, .9, .6];
EdgeColor = [0 0.4470 0.7410];
FaceAlpha = 1;
end
if obj.is_filling
warning('already filling')
return
end
obj.fill_color = FaceColor;
obj.fill_alpha = FaceAlpha;
obj.pen_up();
obj.pen_down(patch(obj.x, obj.y, [1, 1, 1], ...
EdgeColor=EdgeColor, FaceAlpha=0));
obj.is_filling = true;
end
function end_fill(obj)
if ~obj.is_filling
warning('not filling now')
return
end
obj.l(end).FaceColor = obj.fill_color;
obj.l(end).FaceAlpha = obj.fill_alpha;
obj.is_filling = false;
end
function change_icon(obj, filename)
icon = flipud(imread(filename));
obj.im.CData = icon;
obj.im.AlphaData = (255 - double(rgb2gray(icon)))/255;
end
function clear(obj)
obj.x = 0;
obj.y = 0;
delete(obj.ax.Children(2:end));
obj.l = plot(0, 0, 'k');
obj.ht.Matrix = eye(4);
end
end
methods (Access = private)
function animated_step(obj, delta, q, initFcn, updateFcn)
arguments
obj
delta
q
initFcn = @() []
updateFcn = @(~, ~) []
end
dx = delta(1)/obj.n_steps;
dy = delta(2)/obj.n_steps;
dq = q/obj.n_steps;
pause_duration = norm(delta)/obj.speed/obj.speed_reg;
initFcn();
for i = 1:obj.n_steps
updateFcn(dx, dy);
obj.ht.Matrix = makehgtform(...
translate=[obj.x + dx*i, obj.y + dy*i, 0], ...
zrotate=deg2rad(obj.q + dq*i));
pause(pause_duration)
drawnow limitrate
end
obj.x = obj.x + delta(1);
obj.y = obj.y + delta(2);
end
function obj = turn(obj, q)
obj.animated_step([0, 0], q);
obj.q = obj.wrap_angle(obj.q + q, 0);
end
function move(obj, delta)
initFcn = @() [];
updateFcn = @(dx, dy) [];
if ~obj.is_pen_up
initFcn = @() initializeLine();
updateFcn = @(dx, dy) obj.update_end_point(obj.l(end), dx, dy);
end
function initializeLine()
obj.l(end).XData(end+1) = obj.l(end).XData(end);
obj.l(end).YData(end+1) = obj.l(end).YData(end);
end
obj.animated_step(delta, 0, initFcn, updateFcn);
end
function obj = fill(obj, delta)
initFcn = @() initializePatch();
updateFcn = @(dx, dy) obj.update_end_point(obj.l(end), dx, dy);
function initializePatch()
obj.l(end).Vertices(end+1, :) = obj.l(end).Vertices(end, :);
obj.l(end).Faces = 1:size(obj.l(end).Vertices, 1);
end
obj.animated_step(delta, 0, initFcn, updateFcn);
end
end
methods (Static, Access = private)
function update_end_point(l, dx, dy)
l.XData(end) = l.XData(end) + dx;
l.YData(end) = l.YData(end) + dy;
end
function q = wrap_angle(q, min_angle)
q = mod(q - min_angle, 360) + min_angle;
end
end
end
Nice to have - function output argument provide code assist when said function is called
This is a feature which doesn't apear to currently exist, but I think alot of matlab users would like, particularly ones who write alot of custom classes.
Imagine i have a custom class with some properties:
classdef CustomClass < handle
properties
name (1,1) string = "default name"
varOne (1,1) double = 0
end
methods
function obj = CustomClass(name,varOne)
obj.name = name;
obj.VarOne = varOne;
end
end
end
Then imagine I have a function which returns one of these custom class objects:
function [obj] = Calculation(Var1,Var2,name)
arguments (Input)
Var1 (1,1) double
Var2 (1,1) double
end
arguments (Output)
obj (1,1) CustomClass
end
results = Var1 + Var2;
obj = CustomClass(name,result);
end
With this class and this function which returns one of these class objects, I would like the fact that I provided "(1,1) CustomClass" in the output arguemnts block of function "Calculation(Var1,Var2,name)" to trigger code assist automaticaly show me, when writing code that the retuned value from this funciton has properties "name" and "varOne" in the object.
For istance, if I write the following code with this function and the class in the Matlab search path
testObj = Calculation(1,1,"test");
testObj.varOne = 10; %the property "varOne" doesn't apear in code assist when writing this line of code
I would like that the fact function "Calcuation(Var1,Var2,name) has the output arguments block enforcing that this function must return an object of "CustomClass" to make code assist recognise that "testObj" is a "CustomClass" object, just as if testObj was an input argument to another function which had an input argument requiring that "testObj" was a "CustomClass" object.
Maybe this is a feature that may be added to matlab in future releases? (please and thank you LOL)
This is a feature which doesn't apear to currently exist, but I think alot of matlab users would like, particularly ones who write alot of custom classes.
Imagine i have a custom class with some properties:
classdef CustomClass < handle
properties
name (1,1) string = "default name"
varOne (1,1) double = 0
end
methods
function obj = CustomClass(name,varOne)
obj.name = name;
obj.VarOne = varOne;
end
end
end
Then imagine I have a function which returns one of these custom class objects:
function [obj] = Calculation(Var1,Var2,name)
arguments (Input)
Var1 (1,1) double
Var2 (1,1) double
end
arguments (Output)
obj (1,1) CustomClass
end
results = Var1 + Var2;
obj = CustomClass(name,result);
end
With this class and this function which returns one of these class objects, I would like the fact that I provided "(1,1) CustomClass" in the output arguemnts block of function "Calculation(Var1,Var2,name)" to trigger code assist automaticaly show me, when writing code that the retuned value from this funciton has properties "name" and "varOne" in the object.
For istance, if I write the following code with this function and the class in the Matlab search path
testObj = Calculation(1,1,"test");
testObj.varOne = 10; %the property "varOne" doesn't apear in code assist when writing this line of code
I would like that the fact function "Calcuation(Var1,Var2,name) has the output arguments block enforcing that this function must return an object of "CustomClass" to make code assist recognise that "testObj" is a "CustomClass" object, just as if testObj was an input argument to another function which had an input argument requiring that "testObj" was a "CustomClass" object.
Maybe this is a feature that may be added to matlab in future releases? (please and thank you LOL)
I would like to zoom directly on the selected region when using  on my image created with image or imagesc. First of all, I would recommend using image or imagesc and not imshow for this case, see comparison here: Differences between imshow() and image()? However when zooming Stretch-to-Fill behavior happens and I don't want that. Try range zoom to image generated by this code:
on my image created with image or imagesc. First of all, I would recommend using image or imagesc and not imshow for this case, see comparison here: Differences between imshow() and image()? However when zooming Stretch-to-Fill behavior happens and I don't want that. Try range zoom to image generated by this code:
fig = uifigure;
ax = uiaxes(fig);
im = imread("peppers.png");
h = imagesc(im,"Parent",ax);
axis(ax,'tight', 'off')
I can fix that with manualy setting data aspect ratio:
daspect(ax,[1 1 1])
However, I need this code to run automatically after zooming. So I create zoom object and ActionPostCallback which is called everytime after I zoom, see zoom - ActionPostCallback.
z = zoom(ax);
z.ActionPostCallback = @(fig,ax) daspect(ax.Axes,[1 1 1]);
If you need, you can also create ActionPreCallback which is called everytime before I zoom, see zoom - ActionPreCallback.
z.ActionPreCallback = @(fig,ax) daspect(ax.Axes,'auto');
Code written and run in R2025a.
