irf
Generate vector error-correction (VEC) model impulse responses
Syntax
Description
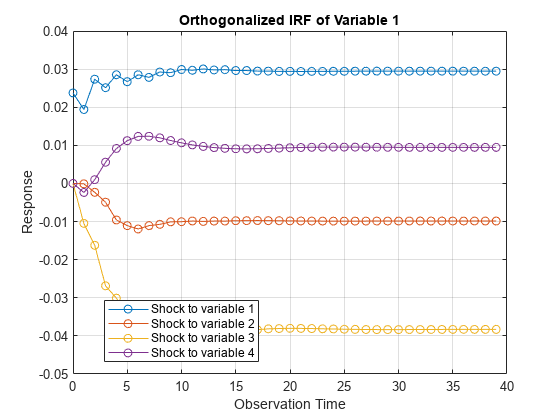
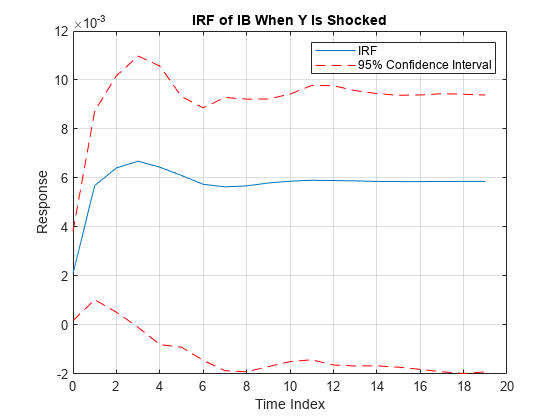
The irf function returns the dynamic response, or the impulse response
function (IRF), to a one-standard-deviation shock to each variable in a VEC(p – 1)
model. A fully specified vecm model object
characterizes the VEC model.
IRFs trace the effects of an innovation shock to one variable on the response of all
variables in the system. In contrast, the forecast error variance decomposition (FEVD)
provides information about the relative importance of each innovation in affecting all
variables in the system. To estimate the FEVD of a VEC model characterized by a
vecm model object, see fevd.
You can supply optional data, such as a presample, as a numeric array, table, or
timetable. However, all specified input data must be the same data type. When the input model
is estimated (returned by estimate), supply
the same data type as the data used to estimate the model. The data type of the outputs
matches the data type of the specified input data.
Response = irf(Mdl)Mdl characterized by a
fully specified vecm model object.
irf shocks variables at time 0, and returns the IRF for
times 0 through 19.
If Mdl is an estimated model (returned by estimate) fit to
a numeric matrix of input response data, this syntax applies.
Response = irf(Mdl,Name,Value)irf(Mdl,NumObs=10,Method="generalized") specifies estimating a
generalized IRF for 10 time points starting at time 0, during which
irf applies the shock.
If Mdl is an estimated model (returned by estimate) fit to
a numeric matrix of input response data, this syntax applies.
[
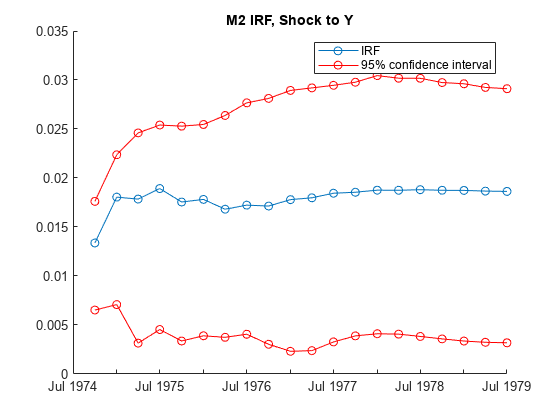
returns numeric arrays of lower Response,Lower,Upper] = irf(___)Lower and upper
Upper 95% confidence bounds for confidence intervals on the true
IRF, for each period and variable in the IRF, using any input argument combination in the
previous syntaxes. By default, irf estimates confidence
bounds by conducting Monte Carlo simulation.
If Mdl is an estimated model fit to a numeric matrix of input
response data, this syntax applies.
If Mdl is a custom vecm model object
(an object not returned by estimate or
modified after estimation), irf can require a sample size for
the simulation SampleSize or presample responses
Y0.
Tbl = irf(___)Tbl containing the IRFs and, optionally, corresponding 95%
confidence bounds, of the response variables that compose the VEC(p –
1) model Mdl. The variables in Tbl correspond to
the variables in the system shocked at time 0. Each variable contains a matrix with
columns corresponding to the IRFs of the variables in the system. (since R2022b)
If you set at least one name-value argument that controls the 95% confidence bounds on
the IRF, Tbl also contains a variable for each of the lower and upper
bounds. For example, Tbl contains confidence bounds when you set the
NumPaths name-value argument.
If Mdl is an estimated model fit to a table or timetable of input
response data, this syntax applies.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
More About
Algorithms
If
Methodis"orthogonalized", then the resulting IRF depends on the order of the variables in the time series model. IfMethodis"generalized", then the resulting IRF is invariant to the order of the variables. Therefore, the two methods generally produce different results.If
Mdl.Covarianceis a diagonal matrix, then the resulting generalized and orthogonalized IRFs are identical. Otherwise, the resulting generalized and orthogonalized IRFs are identical only when the first variable shocks all variables (for example, all else being the same, both methods yield the same value ofResponse(:,1,:)).The predictor data in
XorInSamplerepresents a single path of exogenous multivariate time series. If you specifyXorInSampleand the modelMdlhas a regression component (Mdl.Betais not an empty array),irfapplies the same exogenous data to all paths used for confidence interval estimation.irfconducts a simulation to estimate the confidence boundsLowerandUpperor associated variables inTbl.If you do not specify residuals by supplying
Eor usingInSample,irfconducts a Monte Carlo simulation by following this procedure:Simulate
NumPathsresponse paths of lengthSampleSizefromMdl.Fit
NumPathsmodels that have the same structure asMdlto the simulated response paths. IfMdlcontains a regression component and you specify predictor data by supplyingXor usingInSample, thenirffits theNumPathsmodels to the simulated response paths and the same predictor data (the same predictor data applies to all paths).Estimate
NumPathsIRFs from theNumPathsestimated models.For each time point t = 0,…,
NumObs, estimate the confidence intervals by computing 1 –ConfidenceandConfidencequantiles (upper and lower bounds, respectively).
Otherwise,
irfconducts a nonparametric bootstrap by following this procedure:Resample, with replacement,
SampleSizeresiduals fromEorInSample. Perform this stepNumPathstimes to obtainNumPathspaths.Center each path of bootstrapped residuals.
Filter each path of centered, bootstrapped residuals through
Mdlto obtainNumPathsbootstrapped response paths of lengthSampleSize.Complete steps 2 through 4 of the Monte Carlo simulation, but replace the simulated response paths with the bootstrapped response paths.
References
[1] Hamilton, James D. Time Series Analysis. Princeton, NJ: Princeton University Press, 1994.
[2] Johansen, S. Likelihood-Based Inference in Cointegrated Vector Autoregressive Models. Oxford: Oxford University Press, 1995.
[3] Juselius, K. The Cointegrated VAR Model. Oxford: Oxford University Press, 2006.
[4] Pesaran, H. H., and Y. Shin. "Generalized Impulse Response Analysis in Linear Multivariate Models." Economic Letters. Vol. 58, 1998, pp. 17–29.