Simulate Wild type and Mutant Strains of G Protein Cycle
This example shows how to create and apply a variant to the G protein model of a wild-type strain. The variant represents a parameter value for the G protein model of a mutant strain. Thus, when you simulate the model without applying the variant, you see results for the wild type strain, and when you simulate the model with the variant, you see results for the mutant strain.
The value of the parameter kGd is 0.11 for the wild-type strain and 0.004 for the mutant strain. To represent the mutant strain, you will store an alternate value of 0.004 for the kGd parameter in a variant object, and apply this variant when simulating the model.
Load the gprotein.sbproj project, which includes the variable m1, a SimBiology.Model object.
sbioloadproject gproteinYou can create a variant of the original model by specifying a different parameter value for the kGd parameter of the model. First, add a variant to the m1 model object.
v1 = addvariant(m1,'mutant_strain');Next, add a parameter kGd with a value of 0.004 to the variant object v1.
addcontent(v1,{'parameter','kGd','Value',0.004});Simulate the wild type model.
[t,x,names] = sbiosimulate(m1);
Simulate the mutant strain model by applying the variant.
[tV,xV,names] = sbiosimulate(m1,v1);
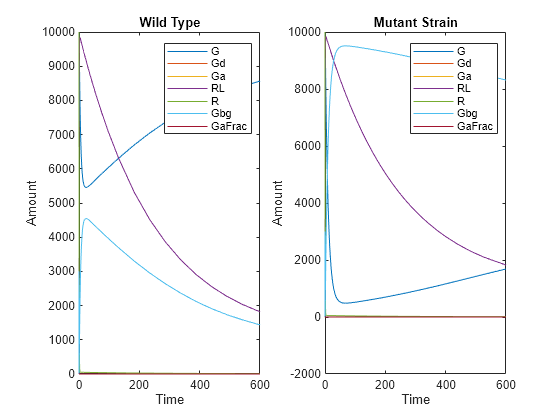
Plot and compare the simulated results.
subplot(1,2,1) plot(t,x); legend(names); xlabel('Time'); ylabel('Amount'); title('Wild Type'); subplot(1,2,2) plot(tV,xV); legend(names); xlabel('Time'); ylabel('Amount'); title('Mutant Strain');
See Also
Variant object | addcontent (variant) | addvariant (model)