View and Run Generated Code by SimBiology Model Builder
You can generate MATLAB® code for modeling actions that you perform in the SimBiology Model Builder app using the Code Capture tool. The tool is useful to learn how to build SimBiology models programmatically or to automate common model building actions at the command line. The generated code shows, for example, how to add model components, configure block properties in the diagram, and change simulation settings programmatically. The following example shows you how to generate code to modify model parameter values and change diagram block properties, save the code as a function file, and run it at the command line.
Load Bioavailability Model
Enter the following command to load the bioavailability model in SimBiology Model Buidler.
openExample('simbio/CodeCaptureSimBiologyModelBuilderExample')
Generate Code for Updating Model Components and Graphical Block Properties
The next sets of instructions show you how to perform the following modeling actions and generate equivalent MATLAB code:
Changing values of some model parameters
Adding a new compartment
Changing the color of all species blocks
Open Code Capture Tool
On the Home tab of the app, select Tools > Code Capture. The app shows the generated code in the Code tab of the Model Tools panel for your modeling actions.
Alternatively, you can expand the Model Tools panel and click Code.

By default, the Code Capture tool continuously generates code for most modeling actions that you do in the app. Specifically, it generates code for model component (object) changes and diagram (graphical) changes. You can choose to generate code for object changes only, diagram changes only, both, or neither (the tool is turned off when you select neither option).

Tip: To capture code for just the actions that you are interested in learning or automating, you can first remove any existing code right before performing those actions by clicking the Clear button of the tool. Alternatively, you can turn off the tool and turn it back on right before the actions.
Change Model Parameter Values
In the Browser panel, scroll down to the Parameters table. Change the values of parameters F (bioavailability of a drug) and ka (total rate of elimination) to 0.5 and 0.7, respectively.
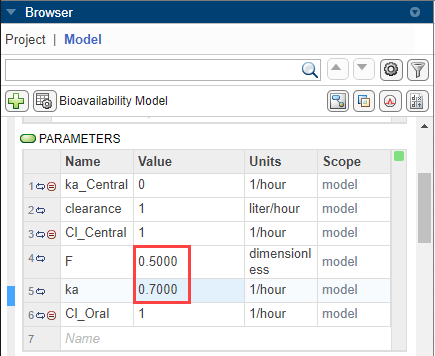
The Code Capture tool shows the following code.
% Query object. p1 = sbioselect(m1, 'Type', 'parameter', 'Name', 'F'); % Configure SimBiology component property. p1.Value = 0.5; % Query object. p2 = sbioselect(m1, 'Type', 'parameter', 'Name', 'ka'); % Configure SimBiology component property. p2.Value = 0.7;
Add New Compartment
Drag and drop a compartment block into the diagram.

Double-click compartment_1 and rename to Brain. Drag and drop a species block into the Brain compartment.
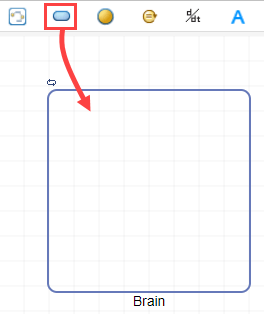
Double-click species_1 and rename to Drug_Brain.
The Code Capture tool shows the following code.
% Add compartment to model. c1 = addcompartment(m1, 'compartment_1'); % Rename SimBiology component. rename(c1, 'Brain'); % Add species to compartment. s1 = addspecies(c1, 'species_1'); % Rename SimBiology component. rename(s1, 'Drug_Brain');
Change Color of Species Block
Click the species Drug_Brain in the diagram. In the Property Editor pane, under the Appearance section, click Face Color and select the green color. Click OK.

The Code Capture tool shows the following code.
% Configure SimBiology block property. simbio.diagram.setBlock(s1, 'FaceColor', [0, 1, 0]);
Tip: For details about this function, see simbio.diagram.setBlock.
Save Generated Code
Click Save in the toolbar of the Code Capture tool.

The app then creates an untitled function file in the MATLAB editor with the generated code in it. Click Save and save the file to your local folder using runcode.m as the file name.
Modify Generated Code
For the purposes of this example, modify the code for the following changes:
Change the parameter values of F and ka to different values.
Add a new compartment called Liver with a species in it.
Change the color of all (instead of one) species in the model to green.
In the runcode.m file, change the values of p1 (parameter F) and p2 (parameter ka).
p1.Value = 0.4; p2.Value = 0.6;
To add a new compartment called Liver with a species named Drug_Liver, update two lines of code to rename the compartment c1 and species s1 as follows.
rename(c1, 'Liver'); rename(s1, 'Drug_Liver');
To change the color of all species to the green color, comment out the setBlock line of code and add two new lines of code as follows.
% simbio.diagram.setBlock(s1, 'FaceColor', [0, 1, 0]); allSpecies = sbioselect(m1, 'Type', 'species'); simbio.diagram.setBlock(allSpecies, 'FaceColor', [0, 1, 0]);
Run Generated Code
To run runcode.m at the command line, you first need to export the corresponding model from the app to the workspace. Go back to the Model Builder app. Select Export > Export Model to MATLAB Workspace. Specify the model name as m1, keep the overwrite check box selected, and click OK.

Keep the app open because the generated code contains graphical changes. Tip: If you have only model component (object) changes, you can optionally close the app after exporting the model to the workspace.
Next, change the current directory to where the runcode.m file is saved. One way to do this is to go back to the Editor window. Right-click the runcode.m tab and select Change Current Folder to.

At the command line, enter:
runcode(m1);
Go back to the Model Builder app to verify the changes you made.
Actions Not Captured
The Code Capture tool does not generate code for the following actions in the app.
Undoing and redoing
Copying and pasting
Dragging or moving a block in the diagram
Diagram layout changes from using the context menu options, such as Layout Method and Layout Diagram
Using the Diagram Alignment Tools to align blocks
Configuring States To Log in Simulation Settings
Updating the Notes section of Property Editor
Using the Model Simulation tool (for details, see Visualize Model Behavior Using Model Simulation Tool)
See Also
SimBiology Model Builder | simbio.diagram.getBlock | simbio.diagram.getLine | simbio.diagram.setBlock | simbio.diagram.setLine | simbio.diagram.splitBlock | simbio.diagram.joinBlock